[#301 | Lost in the Woods] Voting Post
Monday, June 8th, 2026 11:44 pm( List of entries )
Please Note: Because we only have 3 entries this week, there is only a First Place and Runner Up to vote for!
In order to vote, please reply to this post using the form provided. All comments are screened, and entries are listed in the order they were submitted. For your vote to qualify, you must fill out your entire voting card (both spots) in order to be counted. Winner votes are worth 2 points, Runner Up votes are worth 1 point. Meeting the bonus goal on an entry gets an extra point for that submission.
When voting, please copy/paste the ENTRY NUMBER and the FIC TITLE from the list above into the spot you're voting for (this prevents accidentally mis-numbering a vote and casting it for the wrong entry). It should look like this:
First Place: 61. Fic Title Here
Runner Up: 88. Another Fic Title
Please note that you cannot vote for your own entry, and that votes cannot be made anonymously. You do not have to be a member of the community in order to vote, nor have submitted an entry for this week; everyone is welcome to participate in the voting. IP addresses are logged to prevent duplicate voting.
Voting closes Wednesday, June 10 at 9:00PM EST.
Chillicothe was a mess
Monday, June 8th, 2026 11:03 pmI ended up at the Italian restaurant down town. Wasn't planning on it. I checked the hours on the Japanese place in historic downtown. It should have been open. It wasn't. That's okay I wanted to try that Italian anyhow. It was very good.
Somehow this took all day but it was a good one. Right now though, that substernal pressure is back. Can't help but notice more fiberglass has blown up thru my air ducts. I'm beginning to wonder if I have some kind of inflammation going on here as a result (swear to god if this place gives me lung cancer or copd I'm going to become a wrathful spirit)
It's music monday 30 weeks of music. This week's prompt is #29 a song with food or drink in the title
( Notice I know a lot more alcoholic songs than I do food )
here's the whole prompt list
( All under here )
The Personal Cooling Device That Blows Cold Air Rather Than Just Moving It
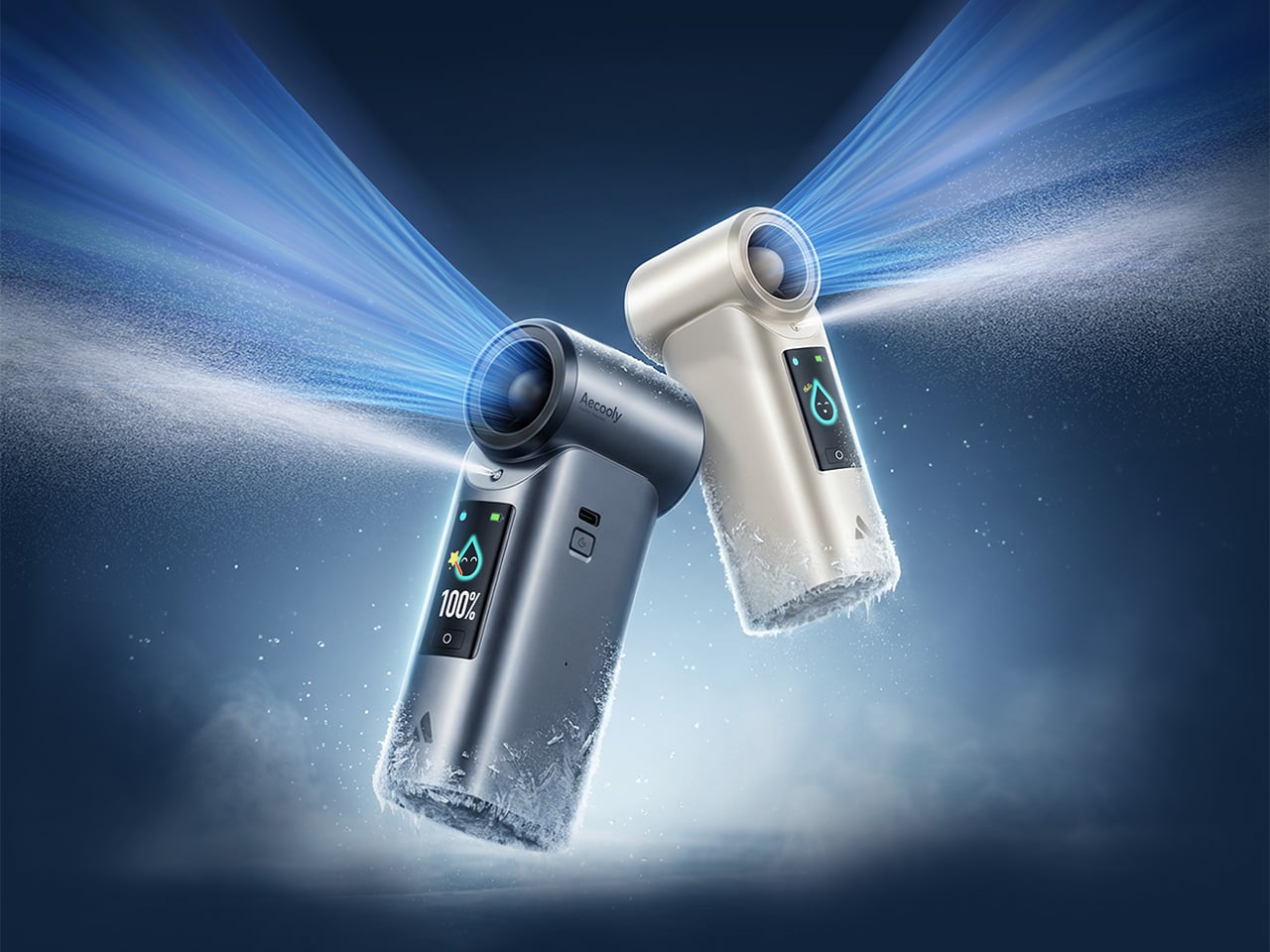
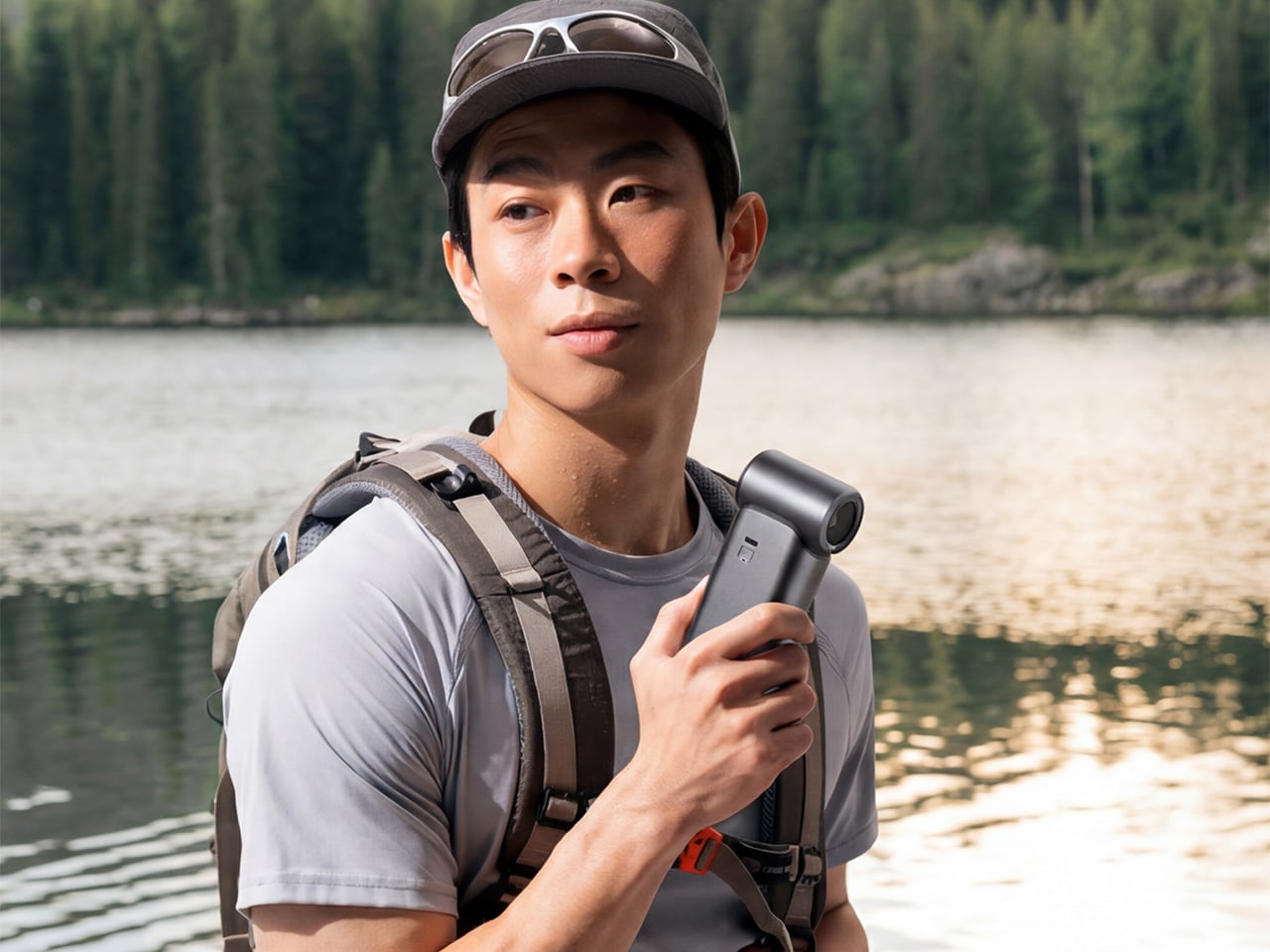
Tuesday, June 9th, 2026 01:46 amHandheld fans have been a summer staple for years, but the basic formula hasn’t changed much. You press a button, blades spin, and air moves. That works fine when it’s mildly warm, but as summers grow hotter and more people spend time outdoors, a fan that simply redistributes hot air starts to feel less like relief and more like a polite gesture against a much bigger problem. And yet for years, that has remained the only option most people know and reach for.
That’s the gap Aecooly is trying to close with the Cold Air Ultra, the brand’s flagship personal cooling device. Rather than simply moving warm air from one side of your face to the other, Aecooly built it around an active cooling system that delivers genuinely cooled airflow, thanks to ultra-fine mist particles that accelerate evaporation to actively draw heat away from the skin. If you’ve ever thought there has to be a fan that actually cools the air, this is that device.
Designer: Aecooly
Click Here to Buy Now: $63.99 $79.99 (20% off, use coupon code “YANKO2026”). Hurry, deal ends in 48 hours!
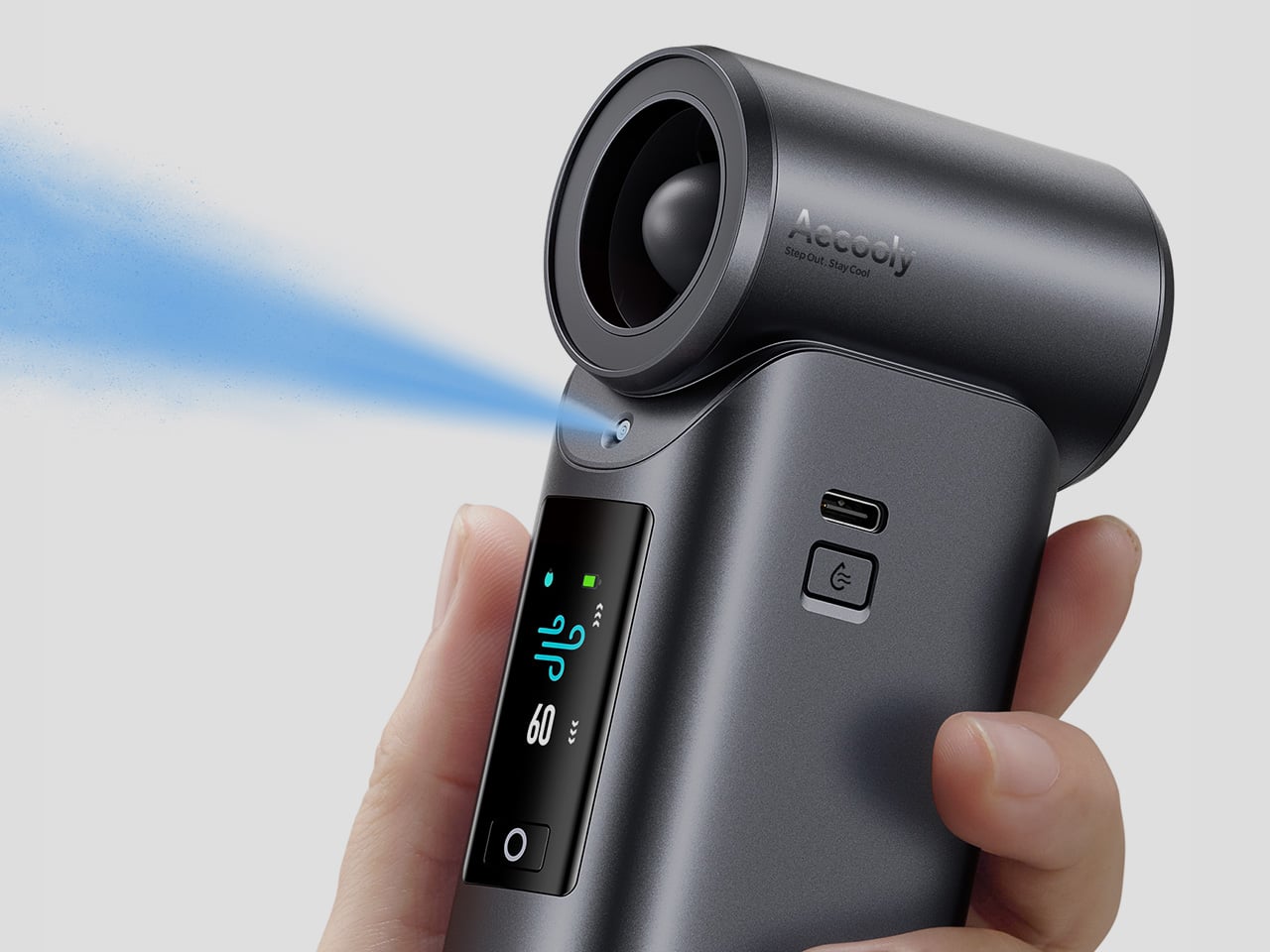
It’s worth noting how the Cold Air Ultra looks, because the design says a lot about what it’s trying to be. The body is compact and upright, with a cylindrical air outlet with a straight, high-pressure duct design, giving it a shape closer to a precision tool than a seasonal gadget and ensuring that the powerful airflow reaches you with zero efficiency loss. Unlike the plastic housing common to most portable fans, the Cold Air Ultra comes in a lightweight body with a premium metallic-inspired finish that’s more resistant to scratches and daily wear, better in hand, and can passively conduct heat away from the motor during extended use.
The cooling technology is where the Cold Air Ultra stands apart. An 70,000 RPM brushless motor drives high-speed air that “breaks through” the sticky sweat layer so the mist can evaporate and pull heat away instantly. It’s this dual action of the wind clearing the path for the mist that makes it possible to reduce skin temperature by up to 18°F (10°C) in just 10 seconds, a noticeably different experience from what you’d get with a standard fan.
What makes the airflow cold rather than simply wet comes down to how the system atomizes water. Rather than using a vibrating membrane to break liquid into droplets, the approach used in most portable evaporative devices, the Cold Air Ultra uses pneumatic atomization: a high-pressure pump forces compressed air through a precision copper nozzle, shearing water into ~20 μm micro-particles at speed. The compression process itself lowers the air temperature before it exits the device, so that what reaches your skin is genuinely cooled airflow, not ambient air with moisture added. The water and air channels are sealed in a patented airtight structure that optimizes flow efficiency and prevents leakage, a design detail that also keeps the electronics fully separated from the water circuit.
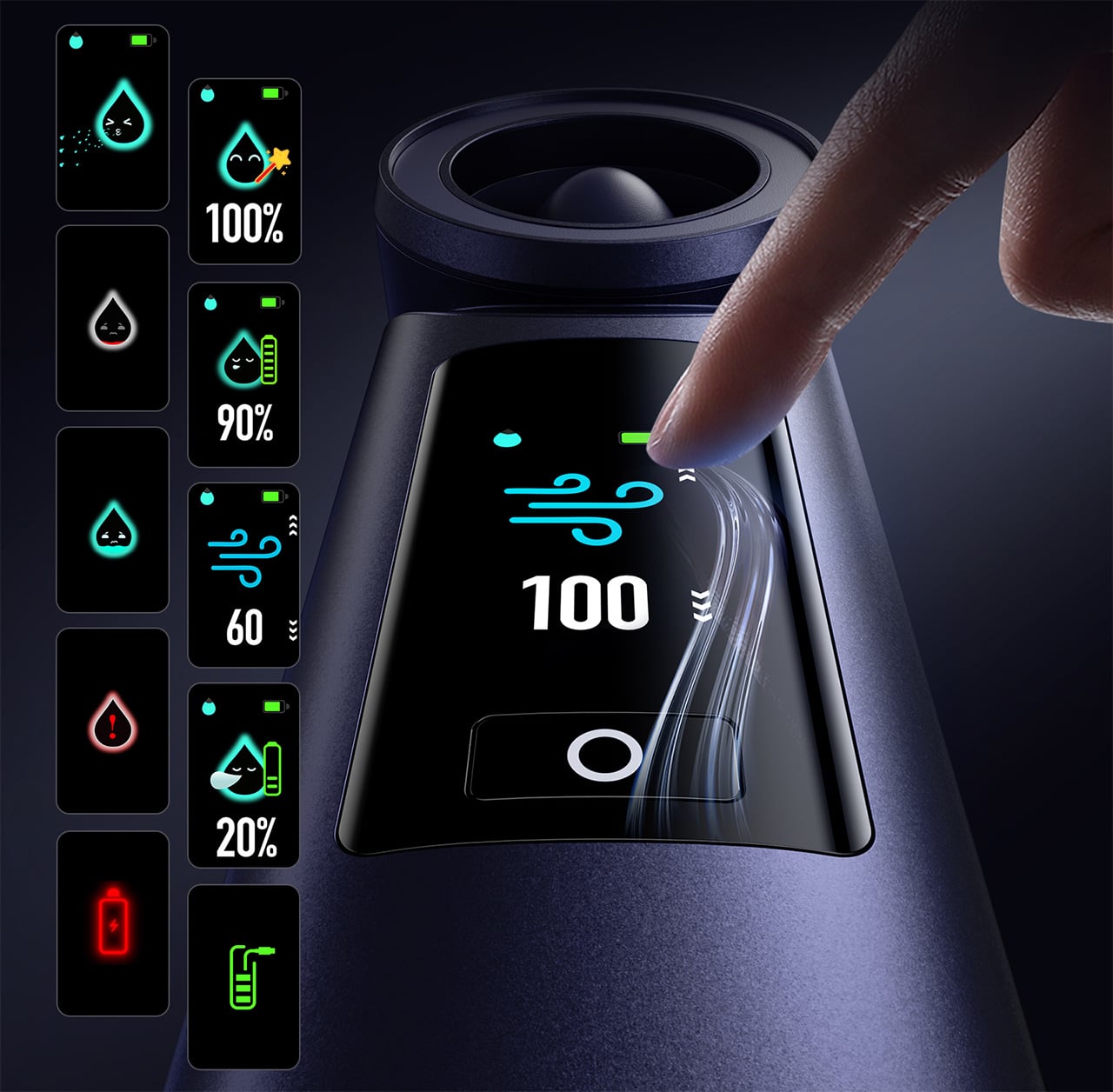
Control is handled through what Aecooly calls the “Little Droplet,” a full-color touchscreen built into the front of the body. This enables fast, intuitive, and precise control, allowing users to swipe through the 100-level settings instantly, which is a much more modern and responsive way to manage airflow compared to traditional fans. And with dynamic icons that display battery level, water level, and mist status in real time, you’re never left guessing what’s left.
Picture stopping mid-hike to cool down, or waiting on a sweltering subway platform with no breeze in sight. A standard fan doesn’t do much in either of those moments beyond moving hot air around. The Cold Air Ultra is built for situations like these, where getting your skin temperature down quickly during a break or a commute actually makes a noticeable difference to how you feel.
Battery life isn’t a compromise here either. The Cold Air Ultra packs a 7,000 mAh cell for up to 10 hours, charges via USB-C in about 2.5 hours, and doubles as a 20W power bank with Quick Charge and Power Delivery support. The magnetic accessory system includes a pointed nozzle and a round nozzle for directing airflow, plus a brush head, each of which can snap on and off without tools. The included lanyard enables hands-free carry, so the device stays accessible during commutes or outdoor use without needing to be held.
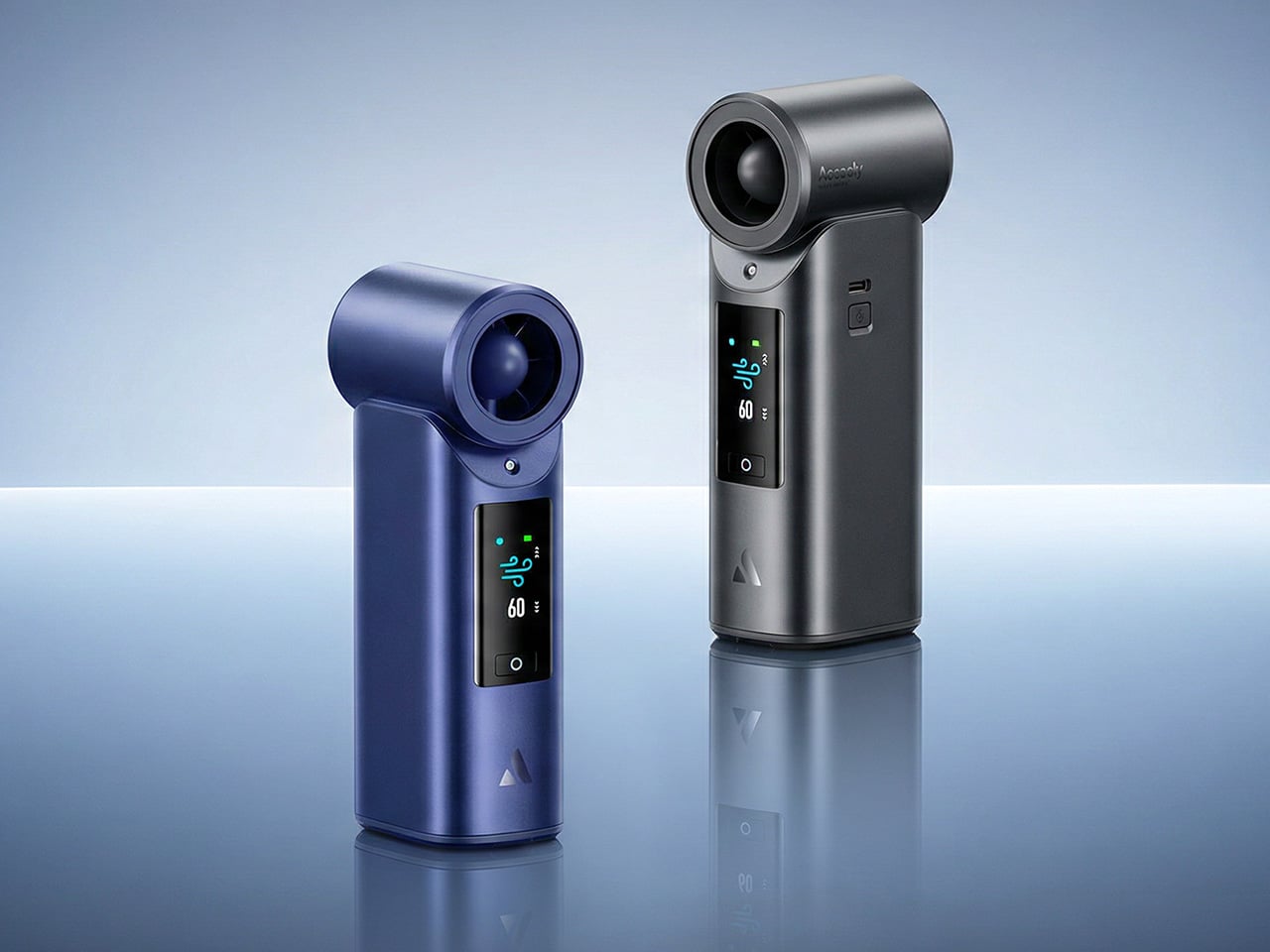
Aecooly says the Cold Air system has received the Red Dot Design Award 2026, a recognition that speaks to its functional engineering as much as its considered form. It’s available in a black and a blue finish, and retails for $79.99. For a device that covers personal cooling, emergency power, and outdoor utility in a single package, the price puts it squarely in premium handheld territory.
The standard Aecooly Cold Air at $31.99 $39.99 (20% off, use coupon code “YANKO2026”) brings the same active cooling concept in a simpler package, with a 4,500 mAh battery and five speed settings. It’s a solid introduction to the concept, but the Cold Air Ultra’s touchscreen, 7,000 mAh battery, 20W power bank output, and magnetic tool system make $79.99 feel less like a premium and more like the smarter spend.
Click Here to Buy Now: $63.99 $79.99 (20% off, use coupon code “YANKO2026”). Hurry, deal ends in 48 hours!
The post The Personal Cooling Device That Blows Cold Air Rather Than Just Moving It first appeared on Yanko Design.
$450 Smart Glasses, 78 Languages, Zero Smartphone Required
Tuesday, June 9th, 2026 12:30 am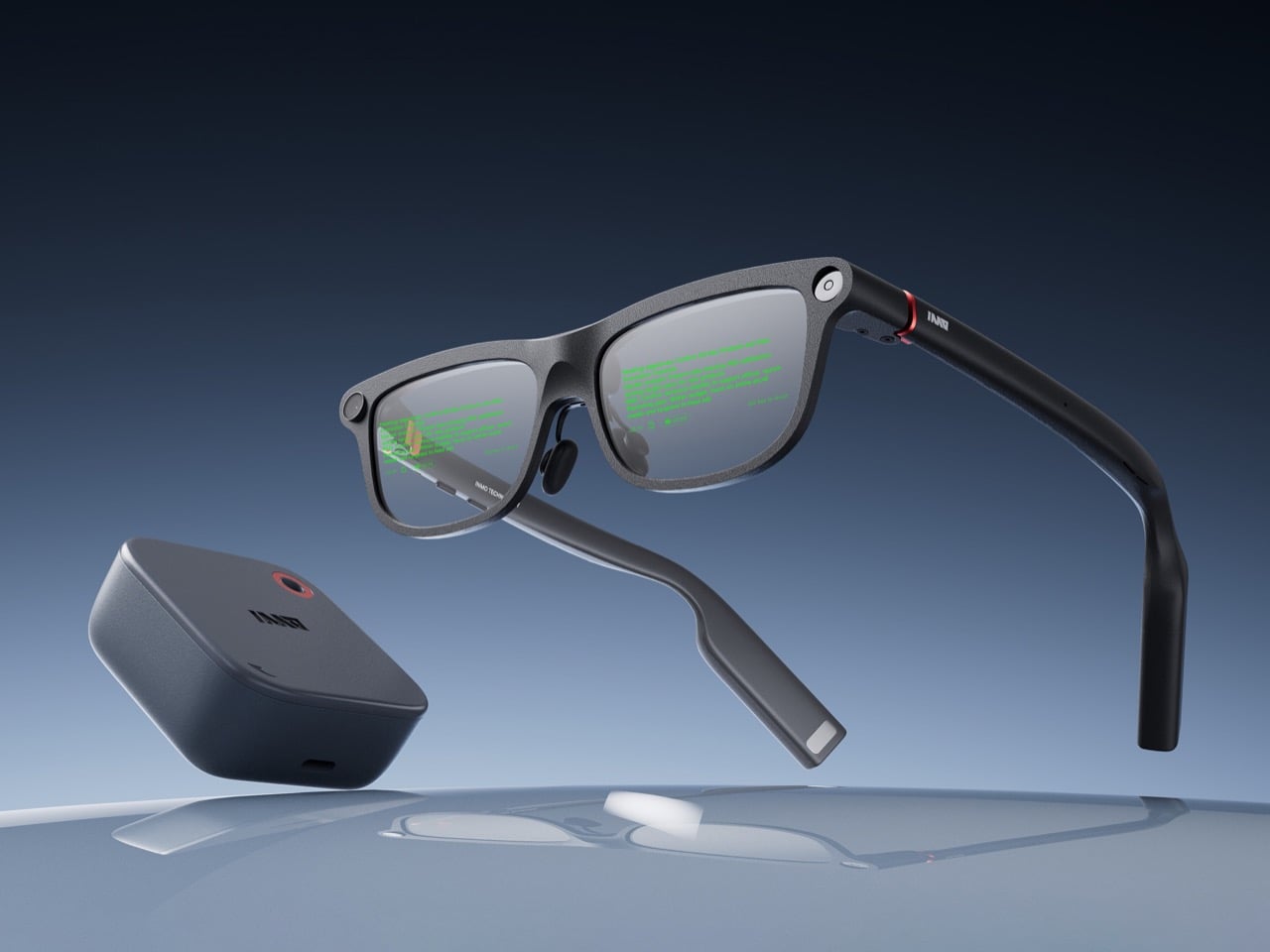
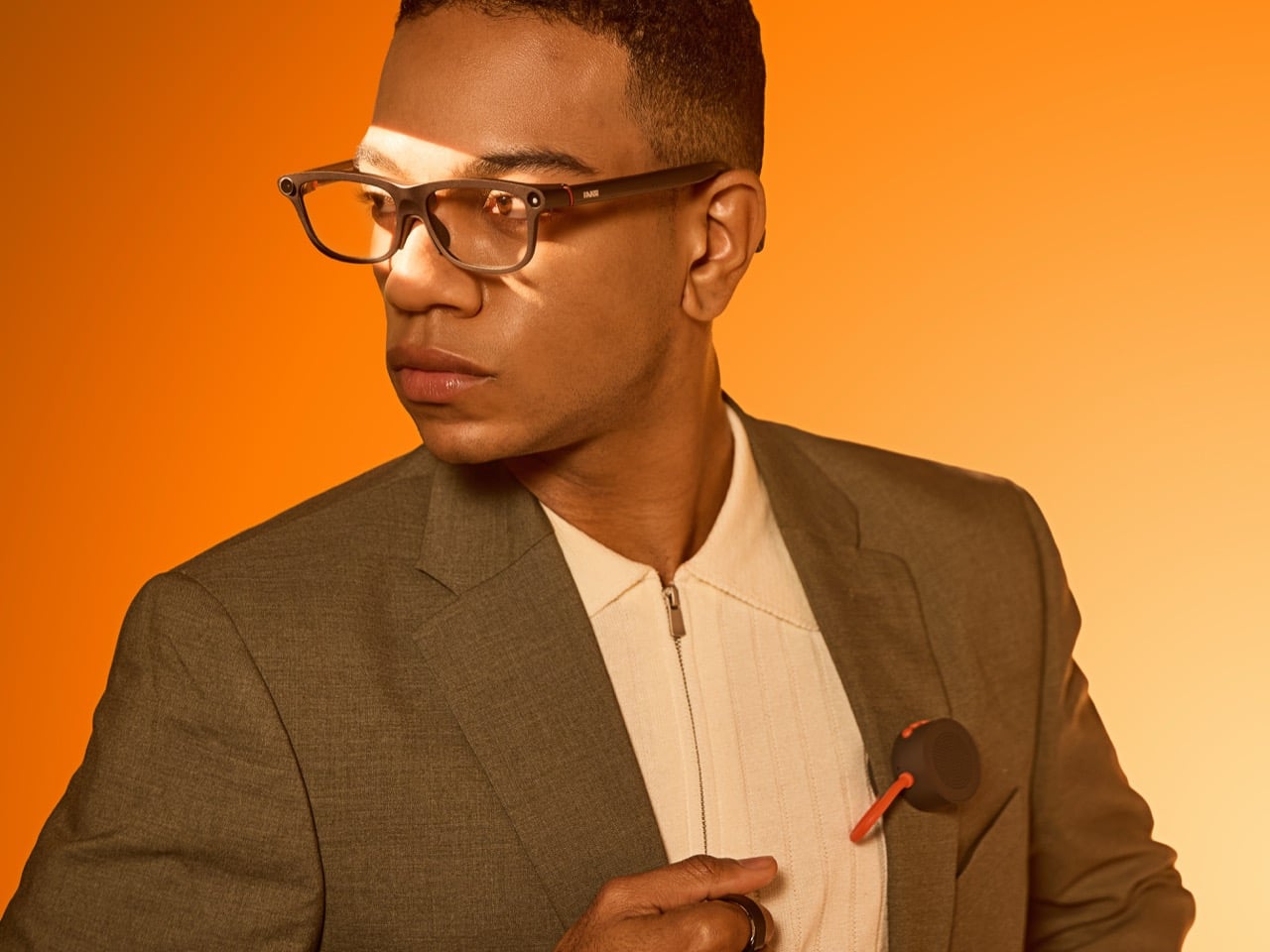
Smart glasses companies love to talk about a grand future, but the strongest case for INMO GO3 arrived in a very ordinary setting: a live presentation. During its appearance at Global Connect in China, an INMO presenter used the glasses’ teleprompter feature while addressing the room, letting the script move with her speaking pace. It was a simple demonstration, slightly funny once the audience noticed the note cards she held as a backup option, and far more memorable than a spec sheet.
That kind of practicality is central to INMO’s broader strategy. The company describes its mission as building glasses people can wear daily, and GO3 reflects that approach with features aimed at frequent, low-friction use. Real-time translation, live transcription, meeting summaries, HERE Maps navigation, and photo translation all point toward the same goal: putting AI and information in front of the eye in a form people might actually keep on their face all day.
Designer: INMO
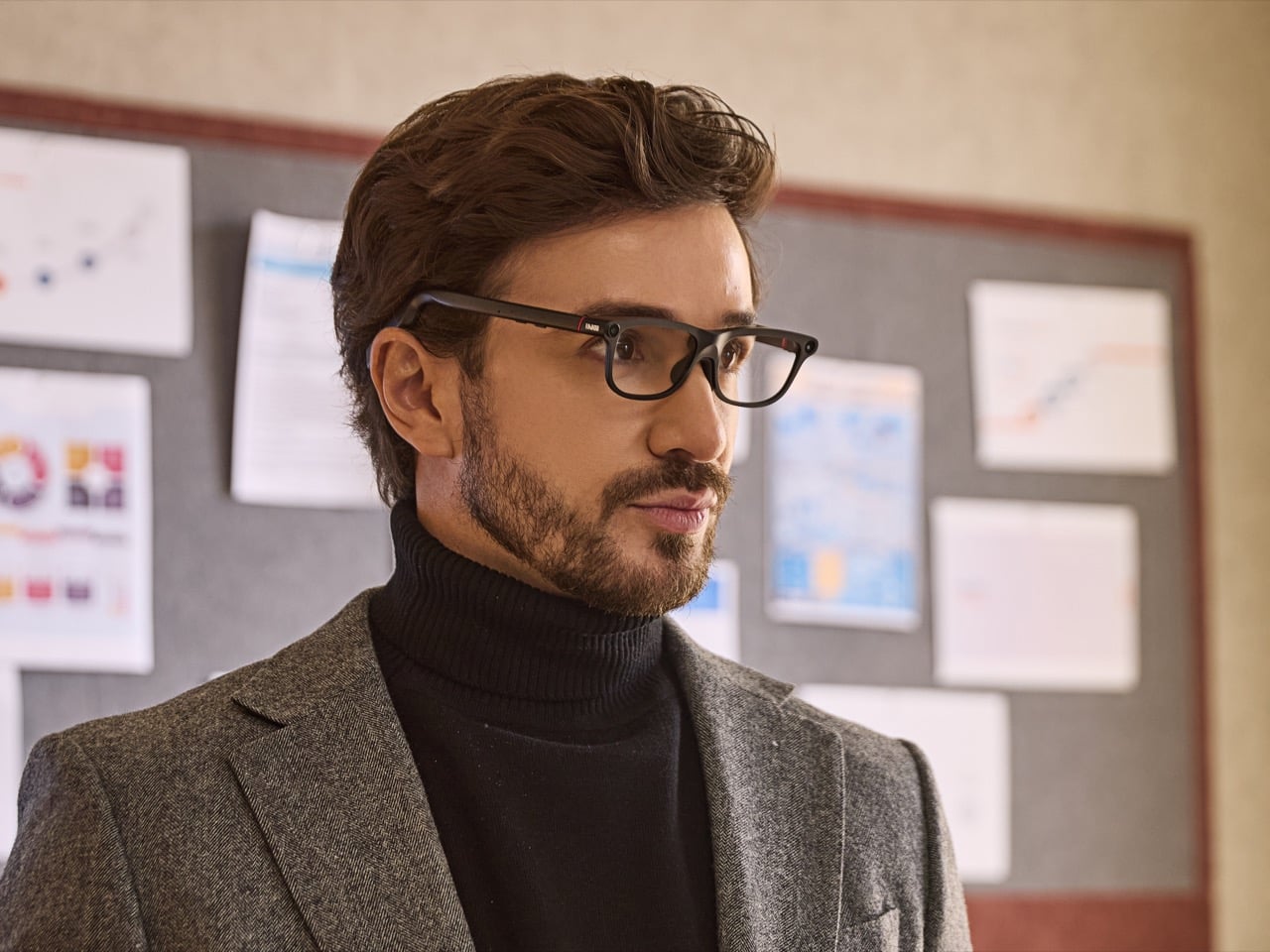

What makes the GO3 feel more meaningful than many smart glasses pitches right now is the kind of display it chooses to be. The category is increasingly pulled toward a model where glasses become another surface for platforms to mediate your attention, observe your behavior, and layer commerce or data collection into the act of seeing. That vision promises convenience, but it also raises the prospect of a device that quietly turns everyday life into a stream of signals for someone else to measure, sort, and monetize.
INMO’s framing, at least from this demo and conversation, points in another direction. The GO3 display feels useful because it serves the wearer in immediate, legible ways. It helps you follow a script. It helps you catch a conversation through live transcription. It helps you understand another language, navigate a route, or pull information from the world through photo translation. The point is not to create a new theater for algorithmic persuasion. The point is to reduce friction between a person and the task in front of them.
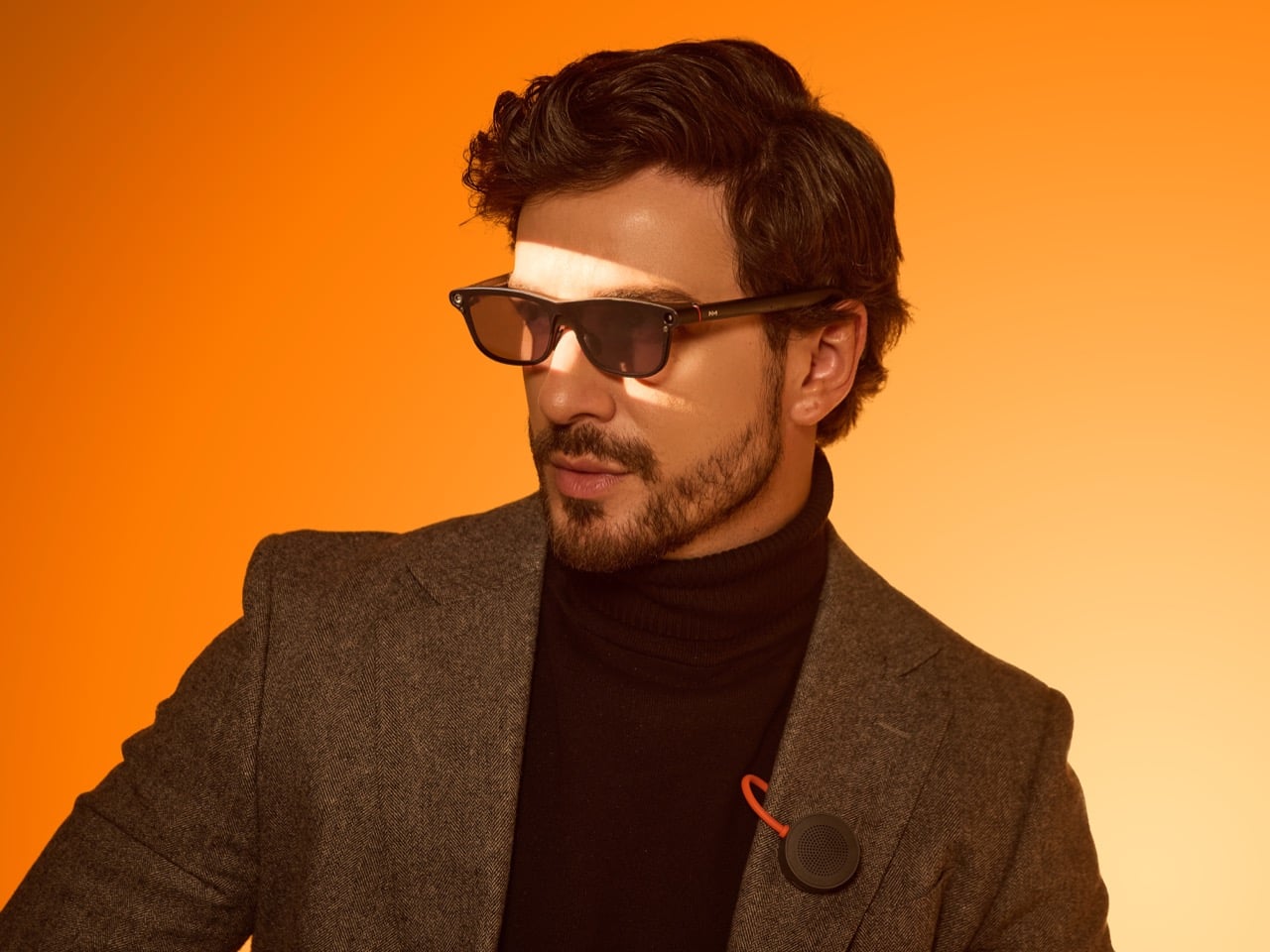

That’s a fairly important distinction because smart glasses will live or die on trust as much as technical ability. People may tolerate a phone screen as a chaotic marketplace of prompts, ads, feeds, and nudges because phones already carry that baggage. Glasses sit closer to the body and closer to perception. They ask for a different kind of acceptance. A product in that position has to prove it deserves to be there, and the most convincing way to do that is by helping with something clear, fast, and human scale.
The GO3 seems to understand that. On paper, its features are varied enough to sound ambitious: standalone real-time translation in 78 languages, AI teleprompting with auto-scroll, meeting summaries, action items, hands-free navigation through HERE Maps, photo translation, prescription support up to 2000 degrees, and a swappable battery system that can be changed in about five seconds. In practice, though, the appeal comes from how these features collapse into ordinary moments. A work presentation. A multilingual conversation. A commute. A quick glance for context instead of a full stop to unlock and consult a phone.

That is why the live teleprompter demo landed so well. It showed the GO3 handling one of the simplest possible tasks, and in doing so, it made a broader case for the category. Smart glasses do not need to begin with spectacle to feel transformative. They can begin with assistance. They can begin with a line of text, quietly placed where you need it, moving at your pace, leaving your hands and attention freer than they were a moment before. Once that works, bigger use cases start to feel plausible.
Some details remain fuzzy, especially around video recording, which was less clearly explained in conversation than photo capture. Any smart glasses company also has to prove that software quality can hold up outside the demo environment, particularly for translation, transcription, and AI-generated summaries. Those are high-value features, but they are also the ones most likely to disappoint if latency, accuracy, or interface design slips. INMO’s been in the business long enough to know that, and to also have a fairly strong grip on a fix.
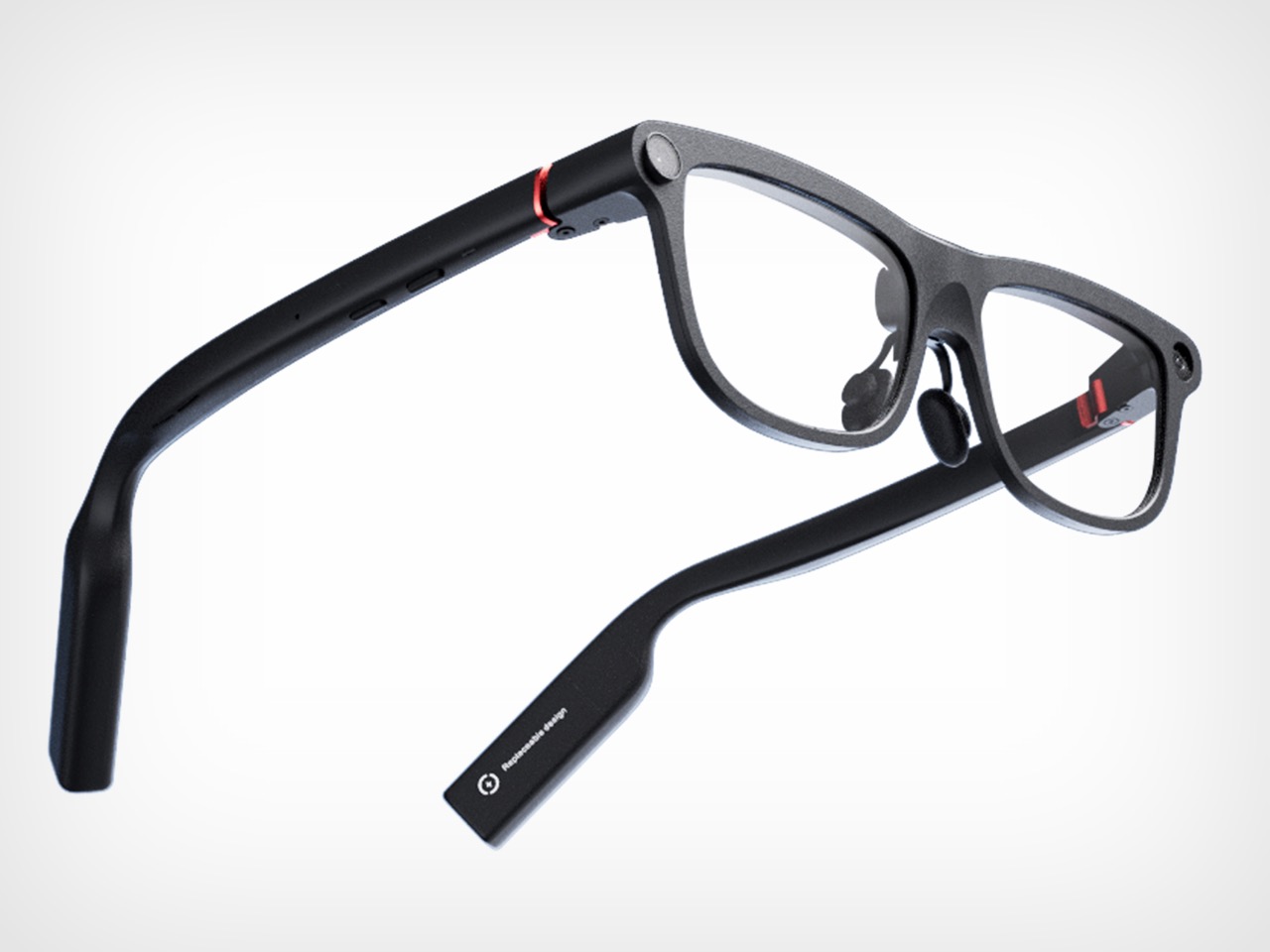

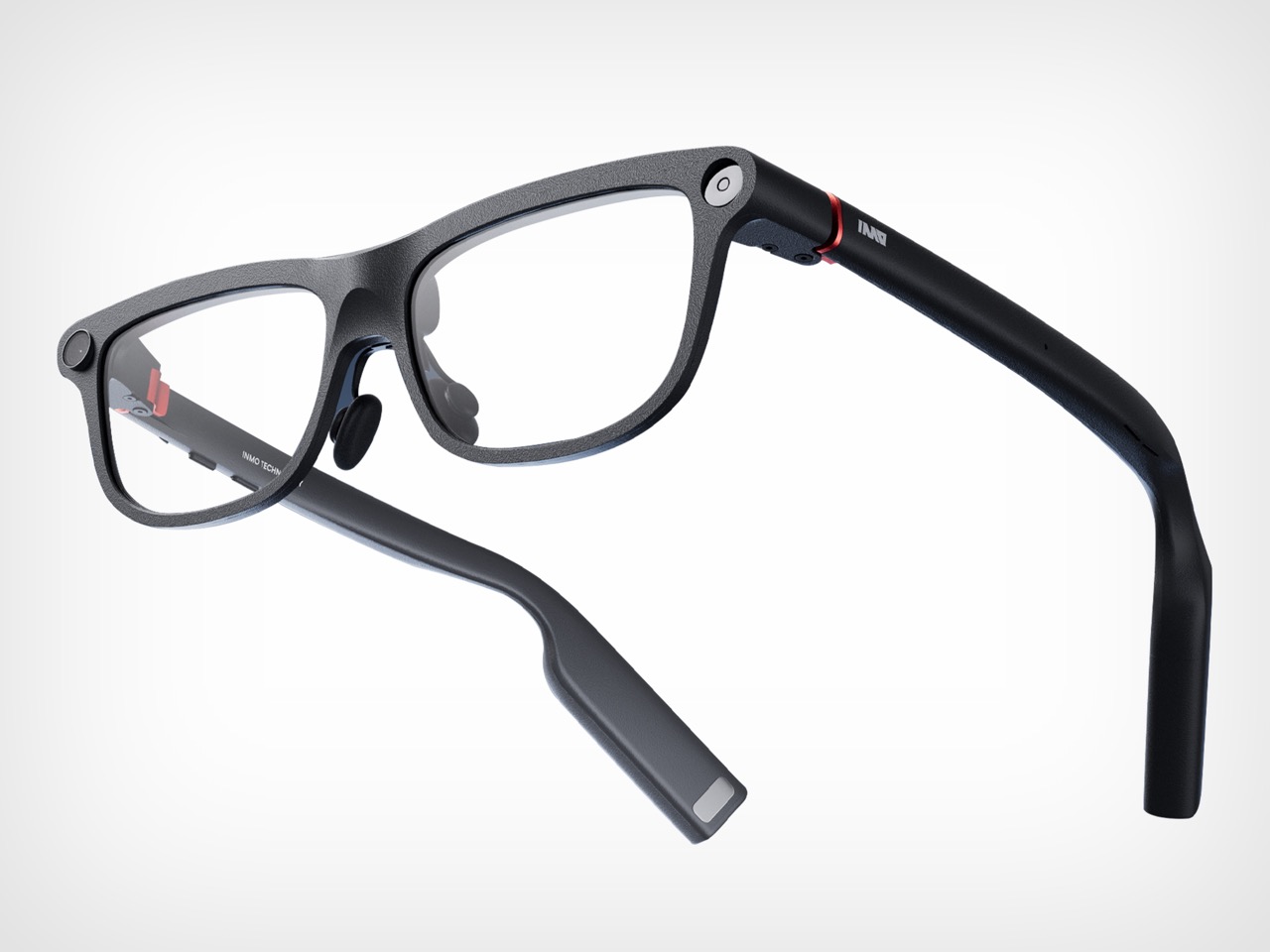
Even with those caveats, INMO’s pitch feels unusually coherent. Founded in 2020, the company says its goal from day one has been to make glasses people will wear every day, and GO3 is the strongest expression of that idea so far. At 58 grams, with prescription support and a battery system designed for long use, it is clearly being shaped around wearability rather than occasional novelty. That design logic gives the product a sense of discipline that many competitors still lack.
The larger vision behind GO3 is that smart glasses will become the next mobile computing platform, eventually taking over as the primary interface for AI. That is a huge claim, and one the industry repeats often. What gives INMO a better argument than most is that it starts from the simple setting rather than the maximal one. If smart glasses are going to matter, they have to prove themselves in the small moments first. GO3 makes that case persuasively. It suggests the future of wearable computing may arrive not through spectacle, but through usefulness that quietly earns its place.

The post $450 Smart Glasses, 78 Languages, Zero Smartphone Required first appeared on Yanko Design.
The Vampire Lestat 1x01 / IWTV 3x01
Monday, June 8th, 2026 09:18 pmThe actors had so much fun, especially Reid. But all of them.
The writers had so much fun with Lestat's voice c. 2025. He's perfectly too much.
The set dressers had so much fun. ( Setting spoilers )
I look forward to ( Character appearance spoilers )
FCC lifts looming deadline for Amazon Leo satellite broadband constellation
Tuesday, June 9th, 2026 12:59 amThe Federal Communications Commission has waived a requirement for Amazon to launch half of its satellite broadband constellation by the end of July, a key regulatory reprieve that buys the tech giant time to get more of its spacecraft into orbit.
Amazon won regulatory approval for the Amazon Leo network in July 2020. The FCC's authorization came with two deadlines. First, Amazon had to launch half of its 3,232 satellites by July 30, 2026, in order to maintain authorization to launch the rest of the network. The regulator gave Amazon a deadline of July 30, 2029, to have all of its first-generation satellites in orbit.
It has been apparent for some time that Amazon would not meet the FCC's requirement to launch half of its satellites—1,616 spacecraft—by the end of next month. Amazon filed an application in January requesting the FCC extend the deadline to July 2028 or waive it altogether. The commission decided on the latter option, removing any time limit for the 50 percent deployment milestone, but keeping the July 2029 deadline in place for the entire constellation.
6/8/2026 The Nature Area
Monday, June 8th, 2026 05:19 pmI wait for them on the bench on the west side of Jewel Lake and we usually sit for a while. There are some very chill Song Sparrows that always come out to be seen, and today several Violet-green Swallows were flying low over the Lake and dipping into the surface, though whether for insects or water or both we could only wonder.
Day 1966: “I didn’t promise anything.”
Monday, June 8th, 2026 05:17 pm
Today in one sentence: Israel and Iran paused direct attacks after trading strikes for the first time since the April ceasefire; Trump denied that he ever promised not to start a war while defending his three-month war with Iran that he launched without congressional approval; Trump abruptly ended a “Meet the Press” interview after Kristen Welker pressed him for evidence supporting his false claims that the 2020 election and California’s primary were “rigged”; the House passed a Ukraine aid and Russia sanctions bill; the Senate passed the $70 billion immigration enforcement bill without restrictions on Trump’s $1.8 billion “anti-weaponization” fund; a federal judge voided Trump’s $100,000 fee on H-1B visa applications; Trump wants his acting director of national intelligence to “start the process” of firing “a lot of people” in the intelligence community; a federal lawsuit seeks to block Trump’s UFC fight on the White House South Lawn arguing the June 14 event violates National Park Service rules, lacked environmental review, and uses federal landmarks for a private, for-profit sports event; and 66% of Americans say a democratically elected government is important to the nation’s identity – down from 80% in 2021.
1/ Israel and Iran paused direct attacks after trading strikes for the first time since the April ceasefire. Trump twice told Israeli Prime Minister Benjamin Netanyahu to stop firing and warned him that Israel could be “on your own very soon.” After Iran fired missiles at Israel in response to Israeli strikes on Hezbollah targets in Beirut, Trump called Netanyahu Sunday night to tell him not to retaliate further. Netanyahu reportedly said Israel had to answer a direct attack and then struck Iranian air defenses and a petrochemical facility, prompting more Iranian missiles toward Israel. Trump then demanded on Truth Social that “Israel and Iran must immediately stop shooting,” and called Netanyahu again Monday morning, and said he told him: “Bibi, you better be careful, or you will be on your own very soon.” Netanyahu later said “the fire on this front has been contained,” but warned Israel would “respond with force” if Iran attacks again. Iran’s armed forces said they were suspending operations, but threatened “more severe and crushing measures” if Israeli strikes continue, including in southern Lebanon. Trump, meanwhile, claimed “final negotiations” with Iran were still proceeding, but Iran’s ambassador to the U.N. said the two sides “have not yet reached a final text.” (New York Times / Axios / Associated Press / NPR / ABC News / NBC News / CNN / Washington Post / Wall Street Journal / Politico / New York Times / Washington Post)
2/ Trump denied that he ever promised not to start a war while defending his three-month war with Iran that he launched without congressional approval. “I didn’t guarantee no war,” Trump said. “I didn’t promise anything.” However, as a candidate, Trump said “I’m not going to start a war. I’m going to stop wars,” and told voters: “I will not send you to fight and die in stupid foreign wars that never end.” Trump insisted Iran “is not an endless war” because “we’ve been doing this for three months,” claimed “the threat is largely over,” and said the U.S. may keep 50,000 troops in the Middle East because “maybe we may use them.” (NPR / New York Times / Politico / The Hill)
3/ Trump abruptly ended a “Meet the Press” interview after Kristen Welker pressed him for evidence supporting his false claims that the 2020 election and California’s primary were “rigged.” Trump claimed California officials were “cheating on the election” because ballots were still being counted days after the primary even though state law allows mail ballots postmarked by Election Day to be accepted for up to a week. “All I have to do is look, and I listen,” Trump said. “But that’s not evidence,” Welker replied. Trump then accused Welker and NBC of being “crooked,” saying “You’re either crooked or you’re stupid” and that “You’re a one-sided crooked network. Sorry. Let’s call it quits because I’ve had enough.” Before removing his microphone and walking off the set, Trump said: “A country can never be great with a dishonest press.” (Washington Post / Politico / BBC / The Hill / Axios)
- The U.S. attorney’s office in Los Angeles opened “multiple election fraud investigations” into California’s primaries and sent a prosecutor to observe ballot processing in Los Angeles County a day after Trump claimed, without evidence, that Democrats were trying to “steal” the races for governor and L.A. mayor. Bill Essayli, a Trump-appointed federal prosecutor, didn’t provide details, but claimed California’s mail-voting system has “serious structural vulnerabilities.” (NBC News / Associated Press / The Hill)
4/ The House passed a Ukraine aid and Russia sanctions bill. 18 Republicans joined Democrats to pass the Ukraine Support Act, 226 to 195, after lawmakers used a rare procedural move to bypass Republican leaders who had kept it off the floor. The bill would authorize up to $8 billion in loans for Ukraine and NATO allies, provide roughly $1.8 billion in military and security aid, while imposing new sanctions on Russia’s energy, finance, and mining sectors, as well as companies and people helping Moscow evade sanctions. The bill, however, faces a difficult path in the Senate and would likely be vetoed if it reached Trump’s desk. (New York Times / Reuters / ABC News / Washington Post / NBC News)
5/ The Senate passed the $70 billion immigration enforcement bill without restrictions on Trump’s $1.8 billion “anti-weaponization” fund, sending the package to the House after Republicans defeated bipartisan attempts to block or limit the taxpayer-funded payout program. The bill, approved 52-47 with Lisa Murkowski the only Republican opposed, would fund ICE and Border Patrol through the rest of Trump’s term using budget reconciliation that bypasses the filibuster. The House is expected to take up the bill next week. Meanwhile, Trump refused to rule out using the fund to compensate Jan. 6 defendants who attacked police, saying he “wouldn’t be inclined” to approve payouts for them, “but I have to see it.” Although acting Attorney General Todd Blanche told lawmakers the administration was “not moving forward with the fund, period,” Trump said “I love the idea” and “if it was up to me, I’d pay them the kind of money that they deserve.” (New York Times / NBC News / The Guardian / NBC News / Politico / ABC News)
6/ A federal judge voided Trump’s $100,000 fee on H-1B visa applications, ruling that the charge was an unlawful tax that Trump had “no power or delegated authority” to impose without Congress. Judge Leo Sorokin said the policy violated the Constitution and the Administrative Procedure Act, and vacated it “in its entirety.” The fee had raised the cost of new H-1B petitions from several thousand dollars to $100,000 for a program used by tech companies, hospitals, universities, and other employers to hire skilled foreign workers. The White House claimed Trump had “clear legal authority” to restrict entry and said the administration was “confident this order will be reversed on appeal,” pointing to a separate December ruling that upheld the fee. (CNN / CNBC / Washington Post / Wall Street Journal / New York Times)
-
A federal judge ordered U.S. Citizenship and Immigration Services to resume processing asylum and immigration applications after striking down Trump administration policies that had frozen decisions for people from 39 travel-ban countries and stopped asylum adjudications worldwide. Chief U.S. District Judge John McConnell said the agency acted without legal authority and “placed the lives of countless individuals on hold — solely by virtue of their countries of birth.” The policies, adopted after an Afghan national was accused of shooting two National Guard members in Washington, left immigrants waiting indefinitely for asylum, work permits, green cards, and citizenship decisions, including lawful permanent residents whose naturalization applications had stopped moving. (Reuters / CNN / The Hill / New York Times)
-
The Trump administration moved to strip citizenship from 17 naturalized Americans accused of hiding crimes, fraud, or false identities when they applied. The Justice Department called it the largest denaturalization push in decades, but the cases still require federal judges to revoke citizenship through a rarely used, difficult court process. (New York Times / CBS News / NOTUS)
7/ Trump wants his acting director of national intelligence to “start the process” of firing “a lot of people” in the intelligence community. Bill Pulte, the director of the Federal Housing Finance Agency, has no national security experience, but can serve for up to 210 days without Senate confirmation. Trump doesn’t plan to nominate Pulte for the role permanently, calling that an advantage because “you’re less shackled.” Trump also said ODNI, which oversees 18 intelligence agencies and units, is “unnecessary and/or too big,” “should maybe even be terminated,” and that Pulte should consider releasing classified documents related to the 2020 election. (Wall Street Journal / The Guardian / Reuters / The Hill)
- The Senate blocked Trump’s effort to extend Section 702 of FISA after seven Republicans joined almost every Democrat to stop a three-year renewal of the warrantless surveillance program, which is set to expire June 12. Section 702 lets U.S. intelligence agencies collect communications of foreign targets overseas, but it also sweeps up Americans communicating with them. (New York Times / CBS News / Wall Street Journal)
8/ A federal lawsuit seeks to block Trump’s UFC fight on the White House South Lawn arguing the June 14 event violates National Park Service rules, lacked environmental review, and uses federal landmarks for a private, for-profit sports event. The Public Integrity Project sued the National Park Service, Interior Department, and Doug Burgum on behalf of two Virginia residents, saying the 92-foot “Claw” structure was built without congressional approval and that ceremonial weigh-ins at the Lincoln Memorial would turn “sacred ground” into a backdrop for “a for-profit cage fight.” The suit also claims UFC, Dana White, and Trump stand to benefit financially, citing VIP and sponsorship packages and Trump’s recent financial disclosure showing a $15,000-to-$50,000 investment in UFC parent company TKO. The White House called the lawsuit “obstructionist, baseless, and dilatory,” saying the event is “no different” from other White House-hosted events. (Politico / NBC News / CNN)
poll/ 66% of Americans say a democratically elected government is important to the nation’s identity – down from 80% in 2021. (Associated Press)
The 2026 midterms are in 148 days; the 2028 presidential election is in 883 days.
- Three years ago today: Day 870: "Blindsided."
- Four years ago today: Day 505: "Transparent."
- Five years ago today: Day 140: "Planned in plain sight."
- Six years ago today: Day 1236: "Now that everything is under perfect control."
- Eight years ago today: Day 505: We have a world to run.
- 9 years ago today: Day 140: No fuzz.
Support today’s essential newsletter and resist the daily shock and awe: Become a member
Subscribe: Get the Daily Update in your inbox for free
Books read, May 2026
Tuesday, June 9th, 2026 12:54 amThe Antiquarian’s Object of Desire, India Holton. Third of the "Love's Academic" series, and I'm glad to say this one felt stronger than its predecessor. It looks like I never posted about that one, so in brief: The Geographer's Map to Romance suffered from a collision between its core trope (the romantic pair are in a marriage of convenience but estranged) and the series pattern of "the characters will spar a lot while secretly being into each other and also sure the other person doesn't reciprocate their feelings." In the first book that worked fine, because the leads were rivals in a contest and started out by thoroughly deceiving one another in pursuit of their goals; it therefore made sense that any signs of romance would fall under suspicion of being just another gambit. But in the second book, it required a degree of emotional stupidity on the part of the characters that I found more grating than charming.
In this third book, the trope is friends-to-lovers, which means the growing warmth between them can be interpreted in that light/suppressed because they don't want to ruin the friendship. Meanwhile, the sparring is because the heroine's job security will be threatened if she's suspected of canoodling with a colleague, so they've agreed to fake-hate. This combination works much better than it did in the previous book. Meanwhile, though I found the magical plot to be slightly muddy in its execution, the ending was entertaining.
I think the series is complete here. Each book stands on its own, though (it's a series in the romance model, where the volumes follow different characters), so you can skip the second one if you want. Me, I think I've had enough of this particular madcap flavor for a while; I overdose on it very easily.
Star*Line 49.2. I've gone ahead and joined the Science Fiction Poetry Association, which means I now have a subscription to their quarterly poetry journal. I don't know that I have a ton to say about it, but poetry was a good match for my short attention span in May!
A Counterfeit Suitor, Darcie Wilde. Another of the Rosalind Thorne Regency mysteries. The mystery in this one did not pull together terribly well for me; there was never a point at which I felt the satisfying "click" of the pieces slotting into place, just "oh, okay, I guess that's what's going on." The personal side was much better, with the heroine's sordid family history rearing its head as a real threat to the life she's built for herself.
At this point I am done with the official Rosalind Thorne series, but I've been told the Useful Woman series is a direct continuation under a different name. So if I want more of these, they're available!
The Bishop’s Tale, Margaret Frazer. As mentioned before, I'm slightly sad that the last couple of books in this series have taken Frevisse out of her nunnery, because one of the things I enjoy here is the view into medieval religious life. However, the usual mystery series consideration applies: you can only have so many murders in one place! Especially when that place is supposed to be cloistered away from the world!
In this case the reason for the departure is very moving, though, and I liked the mystery. It was very obvious to me (as it probably is to many readers) just how the victim actually died -- as opposed to what the characters initially think happened -- but the "who" was less immediately obvious. It also built up to a moment of very effectively understated drama at the end.
The Fallow Year, Margaret Owen. Not actually a novel in the conventional sense, but at over 60K words I'm treating it like one. These are ten connected short stories Owen wrote (and posted to AO3) to cover the year that passes between the second and third books of the Little Thieves trilogy, and what goes on with Vanja and Emeric in that time. I sort of wish I'd known about these stories before I read Holy Terrors, because of course the key events here get described there. If you're invested in the characters, though, it's absolutely worth reading the mini-novel that explores those events in greater detail.
Platform Decay, Martha Wells. New Murderbot! Not my favorite Murderbot, though, I have to admit. It's a perfectly fine extraction mission with good character moments, but at this point I find myself wanting a stronger feeling that some kind of metaplot is approaching culmination, and that's just not what the series is here to do. Murderbot's emotional growth continues, but the external events are much more self-contained, rather than building much on previous installments (though there is a little bit of the latter).
The Water Kingdom: A Secret History of China, Philip Ball, narr. Derek Perkins. This was one of the longer, denser things I started, and the only one I finished this month. I'm not sure audiobook was the best choice: though my familiarity with Chinese names is better than Malagasy ones (cf. last month's post), it's not so excellent that I didn't occasionally lose track of details. Also, while I'm not qualified to judge Perkins' pronunciation, I was irritated by the frequency with which his intonation and pacing announced THIS IS A CHINESE NAME -- he has a tendency to put micro-pauses around them, in a way he doesn't for European names. Possibly that's meant to be an aid for listeners like me, but I found it grating.
The book itself, however, is great! Enough so that I bought a paper copy afterward so I can re-read the sections I'm the most interested in. Ball is comprehensive in his approach to the topic of "water in China": it starts off with information about the hydrology of the region and what its rivers are like, then wanders through the role of water in Chinese philosophy, why it plays such an important practical and symbolic role in politics, historical and modern efforts to control it, how it factors into poetry and art -- you name the angle, there's probably a chapter for it. The result is very interesting both from a "learn more about China" perspective and a "learn more about rivers" perspective.
The Boy’s Tale, Margaret Frazer. Because these are such comfort reads, I ended up reading a second one this month. Yay, we're back at the convent! I had a theory for who the killer was that I quite liked until circumstances pretty obviously spiked that theory, but it would have been in keeping with a pattern I've noticed with Frazer: the killer is rarely A Bad Person Who Deserves Their Punishment. Quite frequently it's someone for whom you're invited to have sympathy -- which does mean that, despite these being comfort reads, I shouldn't pack them too close together. The discovery of the culprit often comes with a side order of feeling bad for how everything fell out, even when I'm enjoying the story.
(originally posted at Swan Tower: https://www.swantower.com/2026/06/08/books-read-may-2026/)
Lake Lewisia #1406
Monday, June 8th, 2026 05:53 pm---
LL#1406
Tests suggest Russian satellites can jam GPS on a continental scale
Monday, June 8th, 2026 09:56 pmRussian satellites have been identified as the cause of mysterious, seconds-long bursts of GPS interference across Europe—a rare example of human-made GPS interference coming from space. But uncertainty still hangs over whether such interference is intentional and if it could be more powerfully weaponized as GPS jamming with continental reach in the future.
The discovery came from an investigation detailed in a June 2 preprint paper by Todd Humphreys and his student Zach Clements at The University of Texas at Austin, along with Argyris Krizise at Stanford University in California. By sifting through public data from ground-based stations with global navigation satellite system (GNSS) receivers, they identified a pattern of high-powered interference lasting less than 10 seconds each time but simultaneously detectable by ground stations across Europe from Norway to Spain to Poland, and even reaching as far west as Greenland and Canada.
By analyzing the ground station data from January 2019 to April 2026, the researchers found 75 days with at least one widespread GNSS interference event overlapping with the GPS L1 frequency band centered on 1575.42 megahertz. That represents the main band used for signal transmission by the US-made GPS satellite constellation and GNSS constellations from other countries.
June 8 - movie time!
Monday, June 8th, 2026 06:44 pmAs possibly a warning to all parents out there, unexpected runaway hit sci-fi movies can imprint themselves on toddlers brains.
Star Wars, retroactively Star Wars: A New Hope, was the first movie that I formed conscious memories around. This was tested at a re-release, when I began telling my dad what was going to happen next. Granted, I was also 'helpfully' translating for Artoo, as I imprinted strongest on him.
I was not quite 2. I know it was not the first I saw in theater; my mother bemoaned the fact that I slept through a re-release of Sleeping Beauty EXCEPT for the dragon.
I do not have the memory of seeing SW in theater myself NOW. But it was family lore that I did know the full plot and character names at that re-release, so I accept it as the first I formed memories of.
Simplify Further’s Goa Tiny Home Fits a Full Life Into 252 Square Feet
Monday, June 8th, 2026 11:30 pm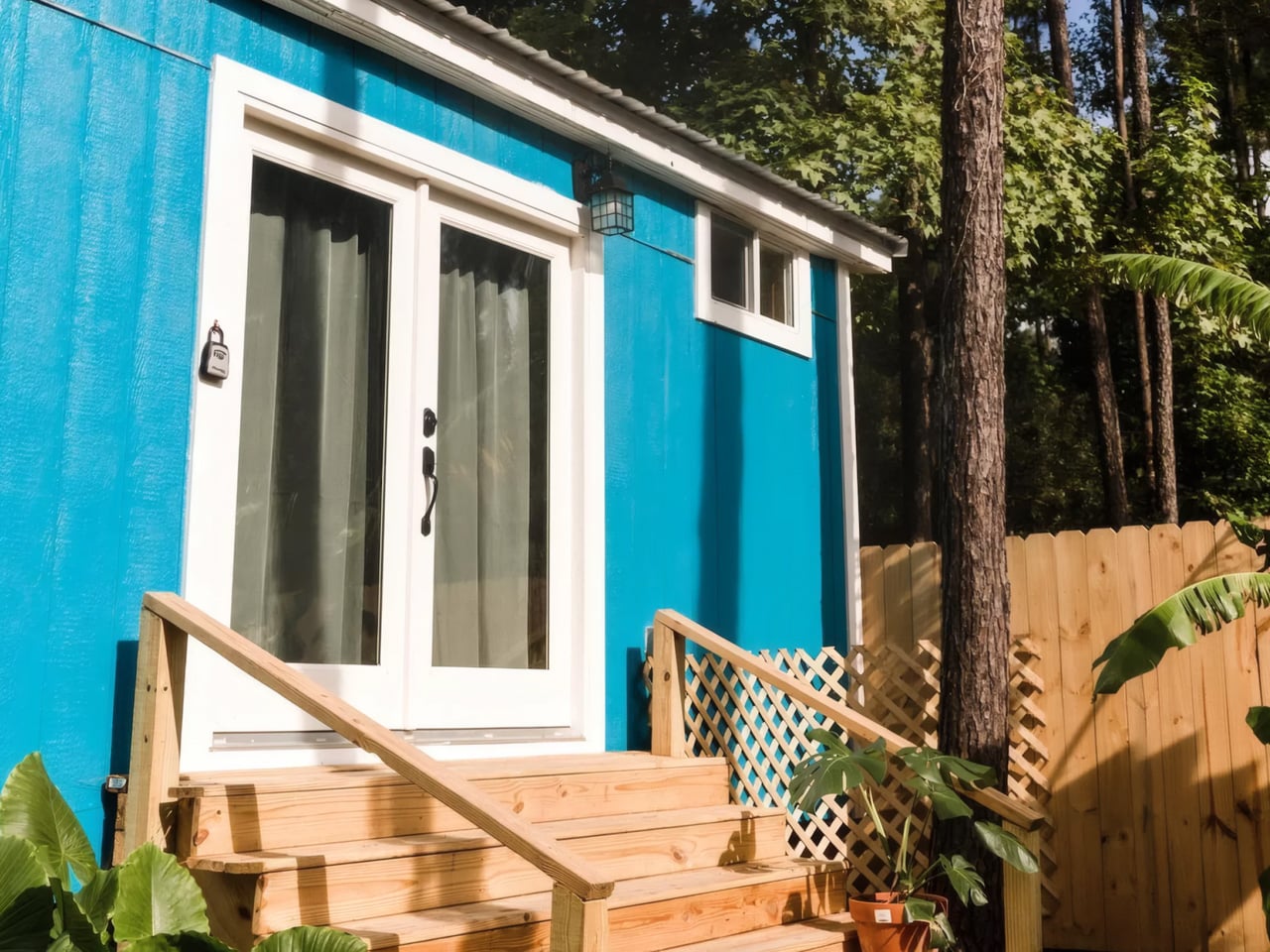
Most tiny homes ask you to give something up. The Goa by Simplify Further Tiny Homes is built around the idea that you shouldn’t have to. It’s a 24 x 8-foot home on wheels designed for people who want to genuinely live small, not just survive it. At 252 square feet, the Goa is built to sleep four to five people, which already tells you something about how thoughtfully the space has been planned.
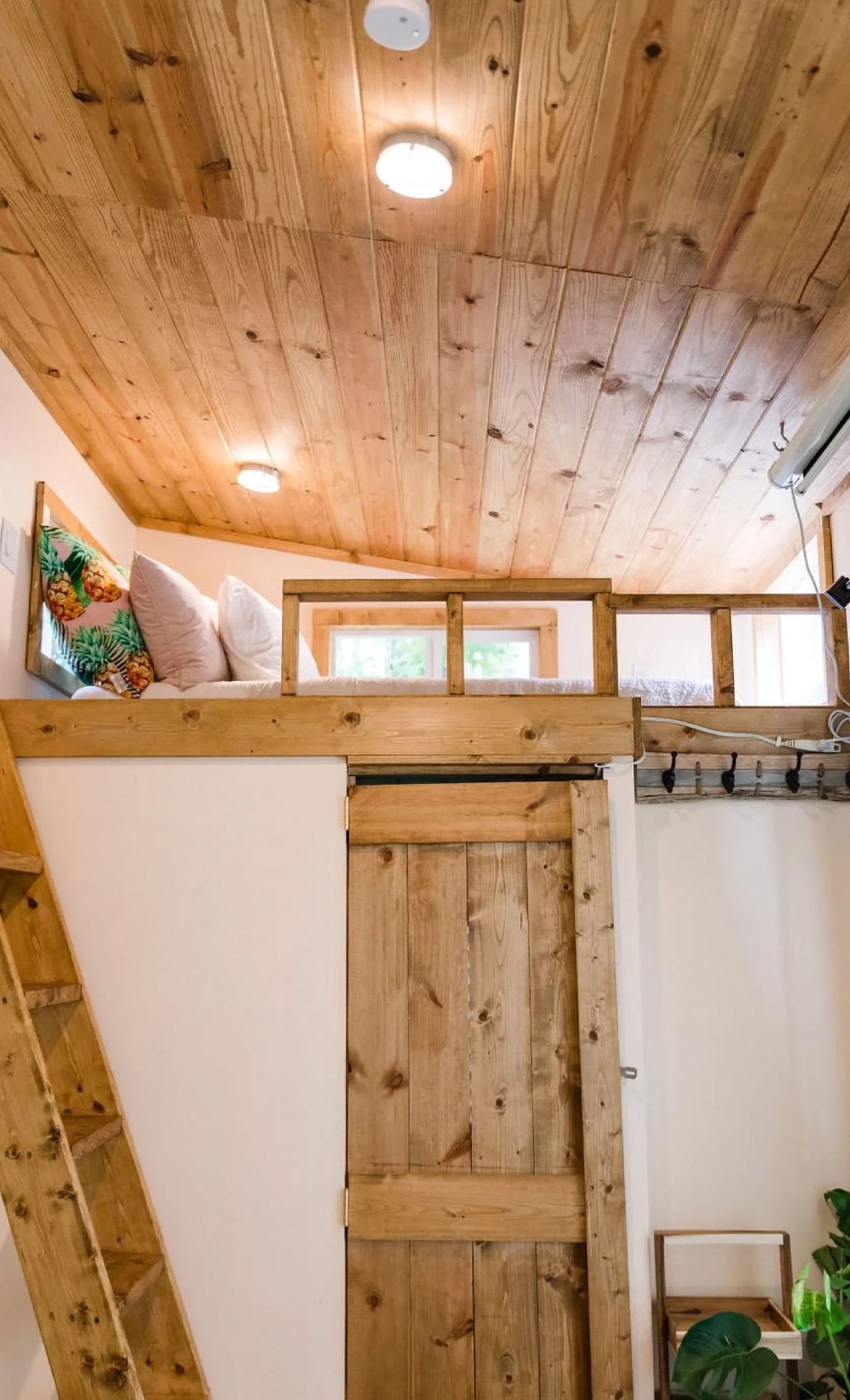
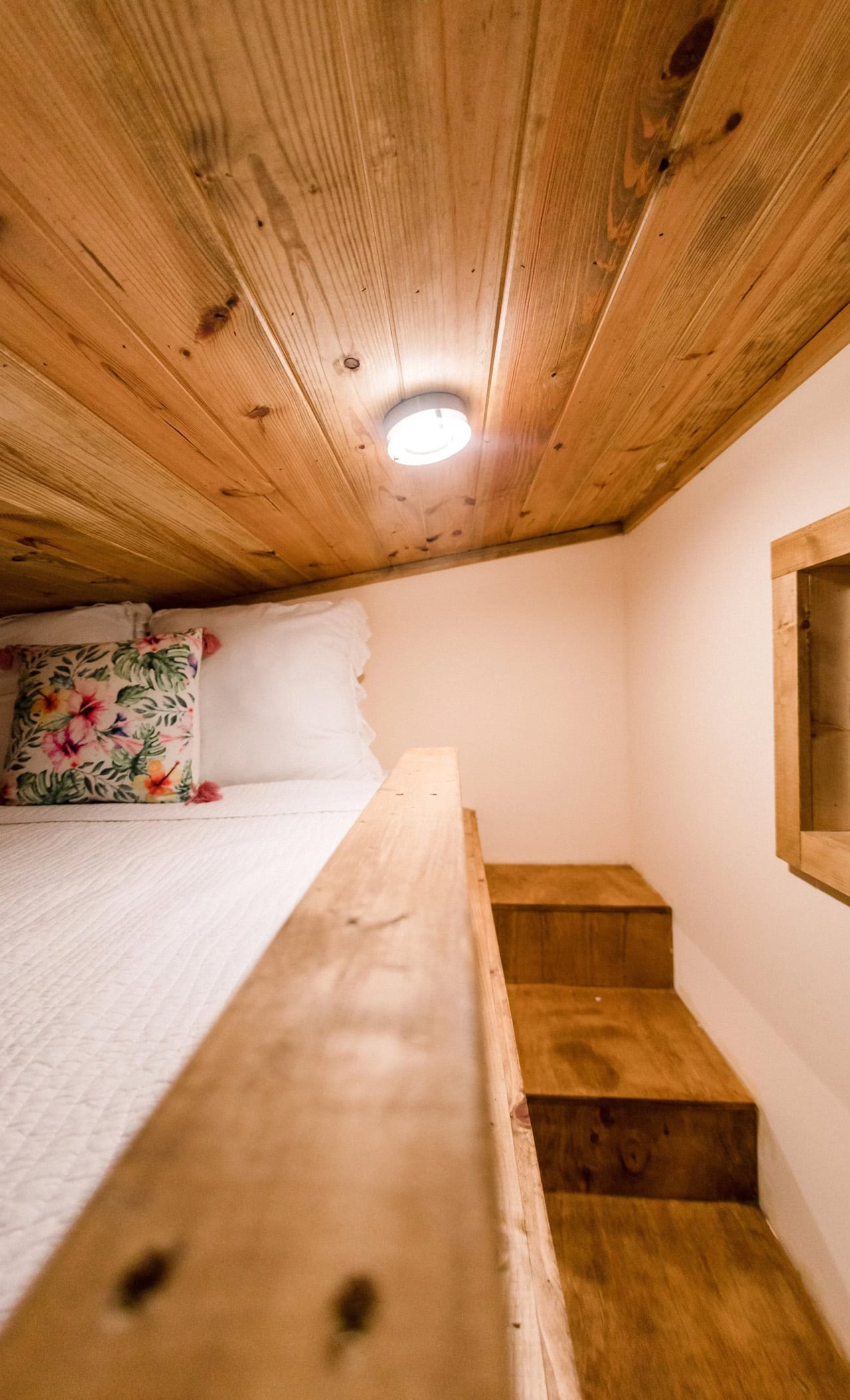
Two sleeping lofts — one measuring 7×8 feet and another at 7×5 feet — sit overhead, leaving a loft height clearance of 36 inches at the low side and 6 feet 4 inches of headroom beneath them. It’s a layout that stacks the private spaces upward and reserves the ground level for living, cooking, and everything in between.
Designer: Simplify Further Tiny Homes
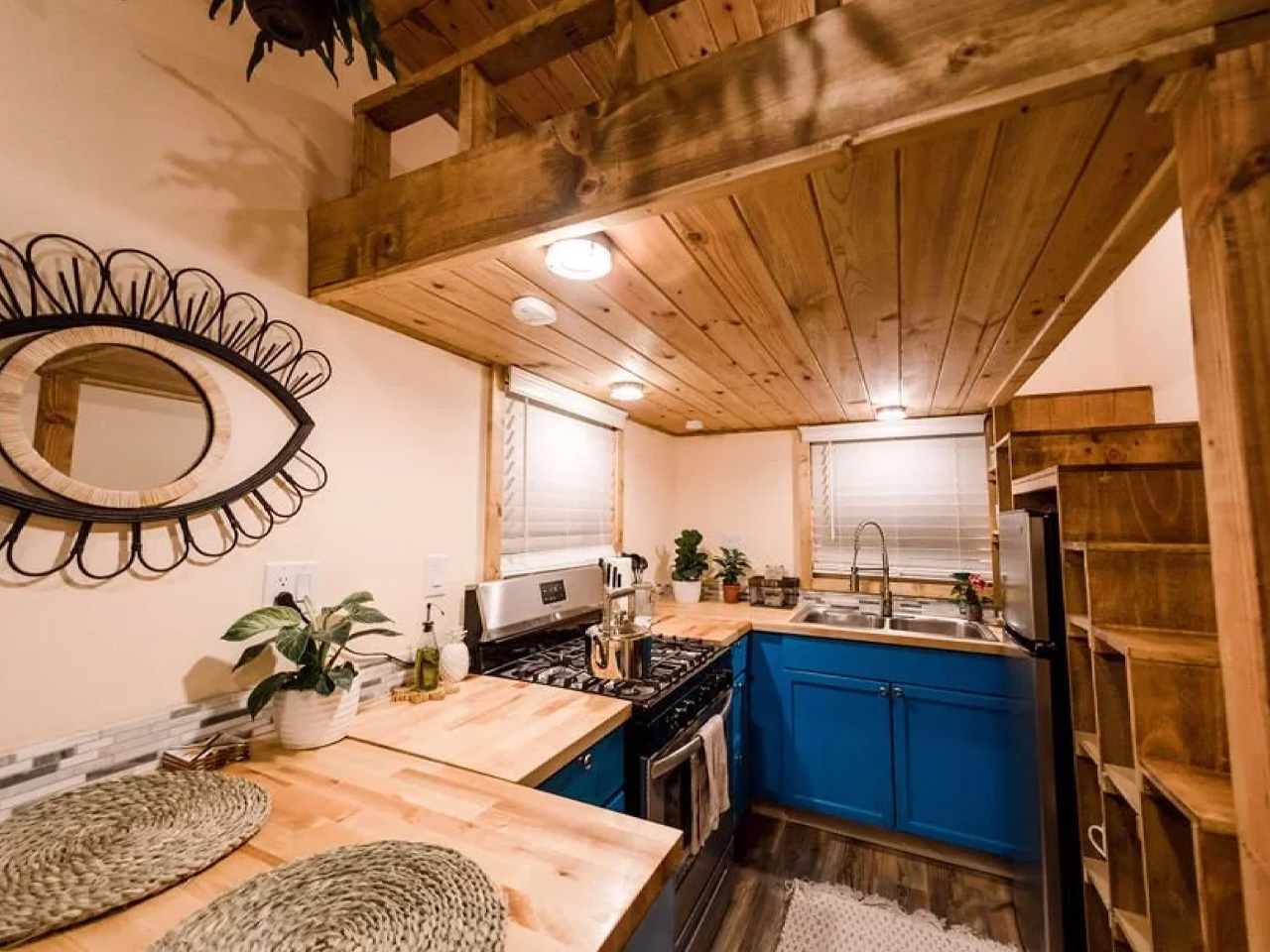
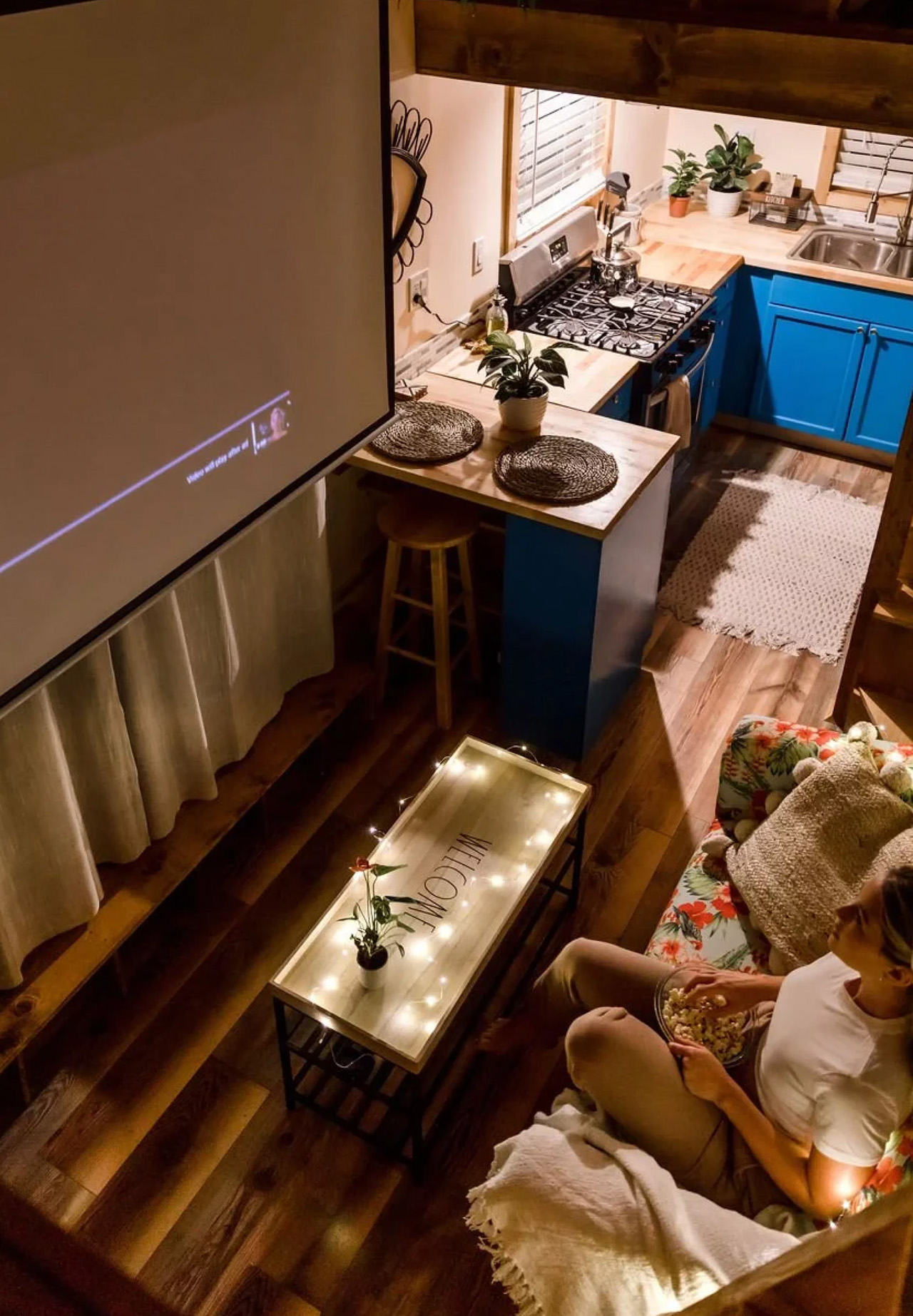


The kitchen is the centerpiece of the Goa, and Simplify Further leans into that fully. A U-shaped layout tucks beneath one of the sleeping lofts, fitted with a four-burner electric range, a 7.1 cubic foot refrigerator, and generous built-in storage — including more tucked beneath the staircase that leads to the loft. It’s a kitchen that actually invites you to cook, not just reheat. A small dining table and seating area sit nearby, keeping the social flow between the kitchen and living room easy and natural.
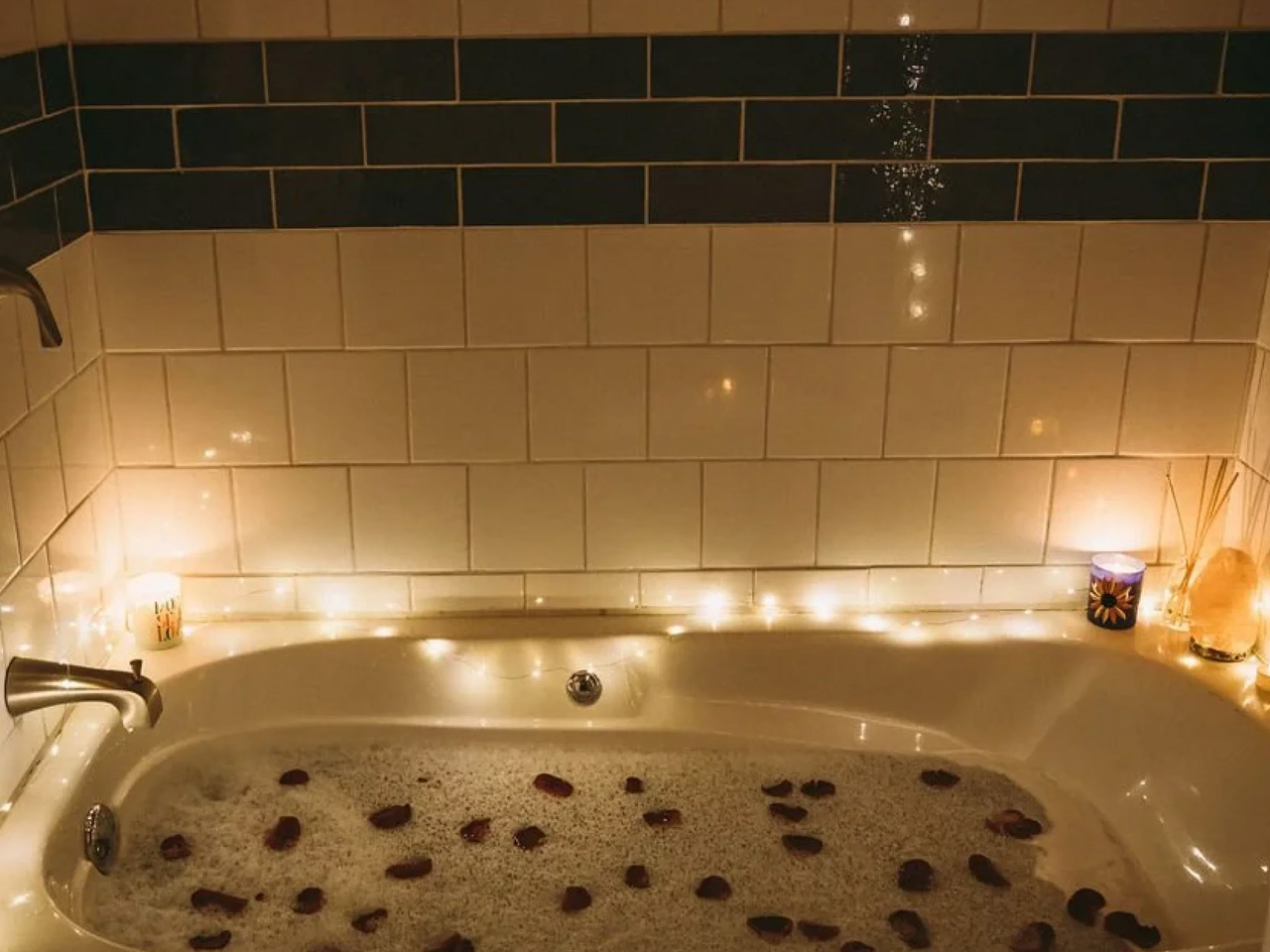
The bathroom is full-sized — a detail that shouldn’t feel remarkable but often does in homes this compact. Buyers can opt for a full-size bathtub or a 36-inch shower with additional storage, depending on how they want to use the space. A washer/dryer combo is also included as standard, which rounds out the Goa as a proper full-time residence rather than an extended camping experience.
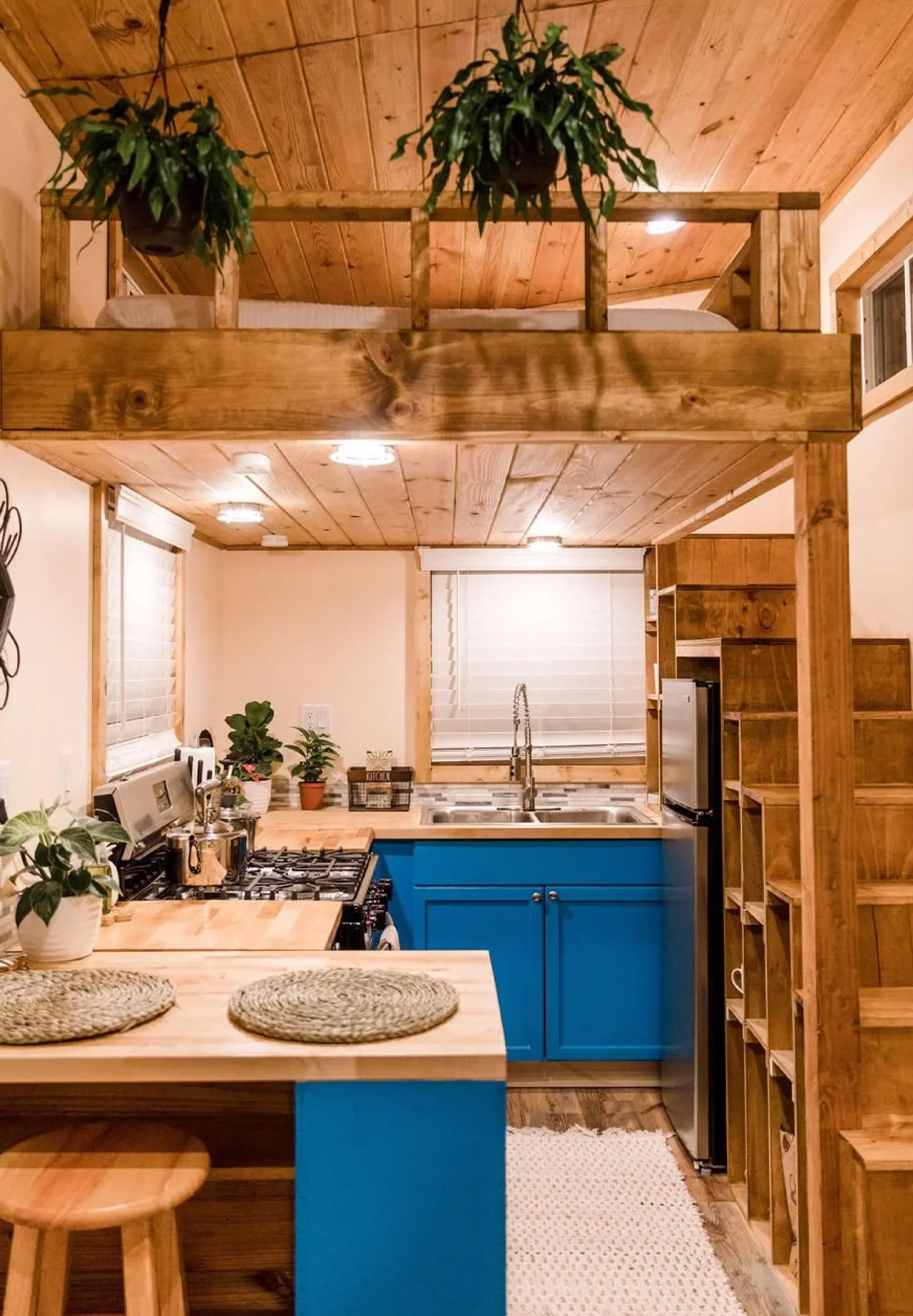
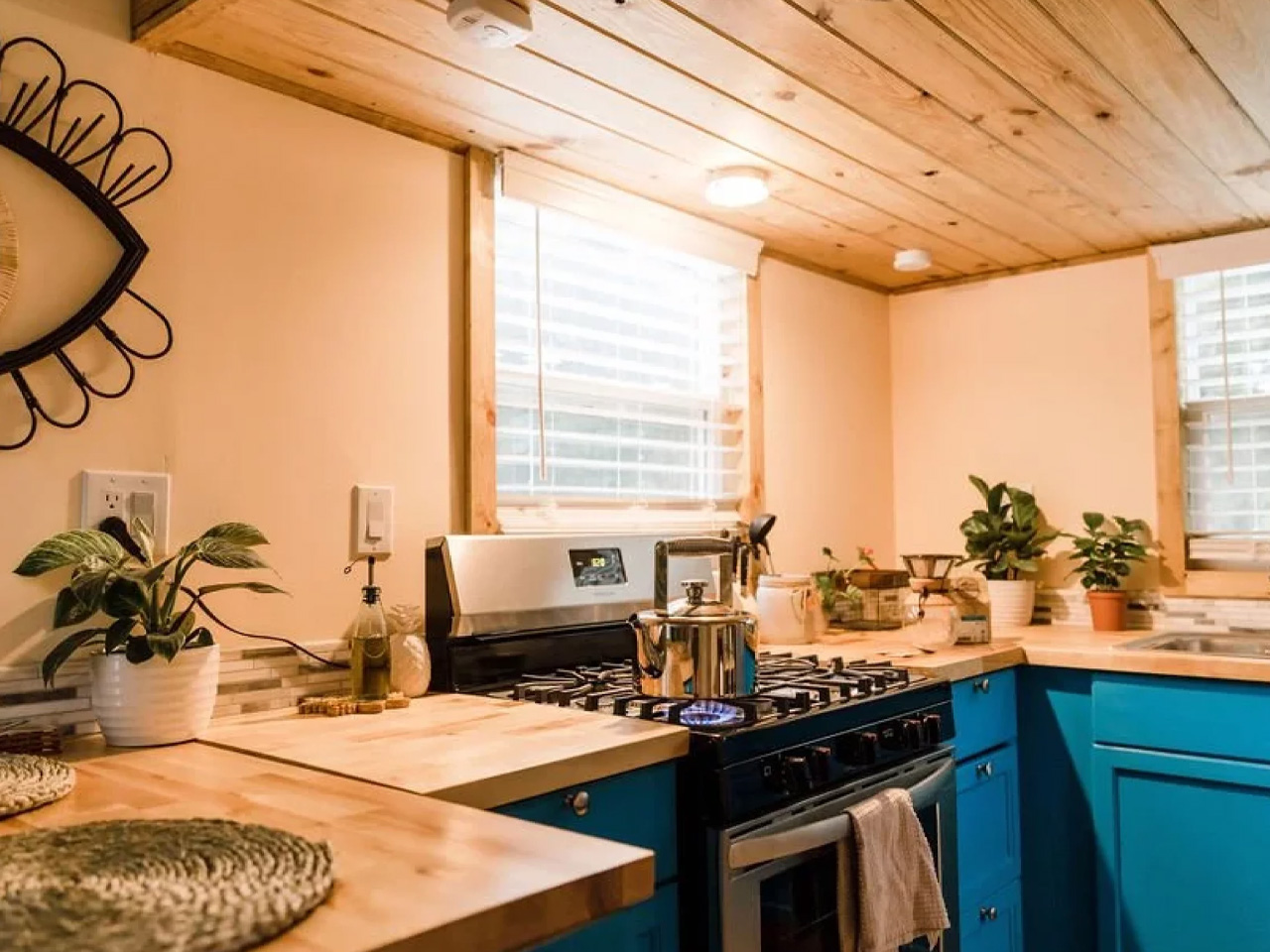


Finish-wise, the interior is dressed in drywall, pine tongue-and-groove ceilings, and vinyl flooring — warm without trying too hard. Upgrade options include shiplap interior walls and furnishings for those who want to move in without lifting a finger beyond signing a check.
The Goa rolls on a hand-built chassis with double axles rated at 7,500 pounds each, trailer brakes, and DOT-approved highway lighting. It carries NOAH certification as an RV and can also be built to satisfy IRC Appendix AQ standards by request. Starting at $65,000, the Goa lands as one of the more compelling full-time tiny home options on the market — a house that earns its footprint rather than apologizing for it.



The post Simplify Further’s Goa Tiny Home Fits a Full Life Into 252 Square Feet first appeared on Yanko Design.
Crocs Just Entered the Mushroom Kingdom and We’re Into It
Monday, June 8th, 2026 10:30 pm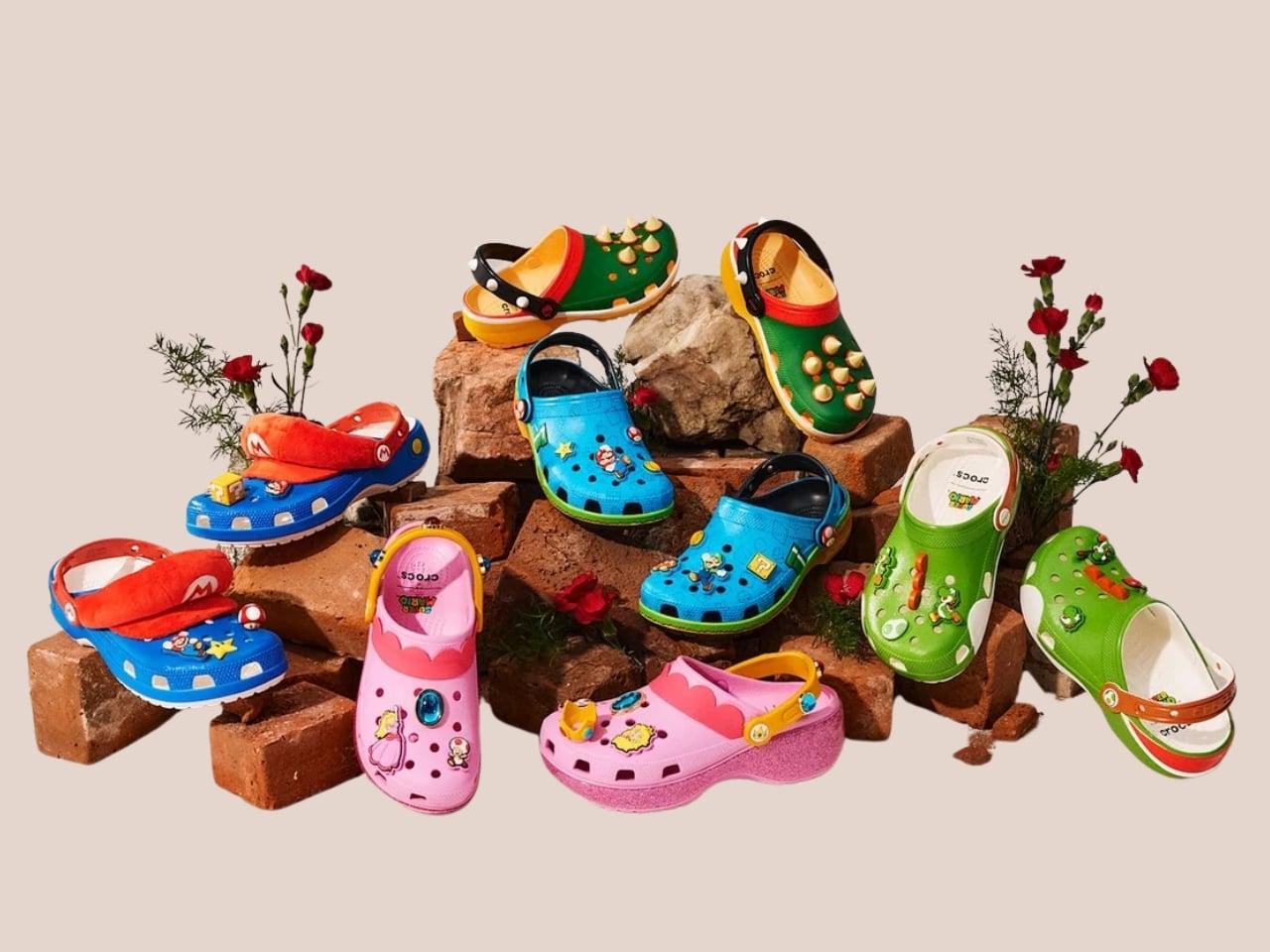
Crocs and Nintendo are not brands you’d typically put in the same sentence as “high-concept design collaboration.” And yet here we are, with the Crocs x Super Mario collection dropping July 15, 2026, looking far better than the premise suggests it should.
The announcement landed earlier this week, and the first reaction — at least my first reaction — was that resigned nod you give when two things that were always meant to go together finally figure it out. Crocs has been quietly rebuilding its cultural credibility for years through a series of increasingly clever partnerships, and Super Mario is the one IP on the planet that genuinely belongs to everyone. People who grew up with a Game Boy, adults who still boot up Mario Kart on a Friday night, and people who have never touched a controller but know exactly who Mario is. That kind of universal ownership is rare, and landing it on a foam clog might be the smartest move either brand has made this year.
Designer: Crocs x Nintendo
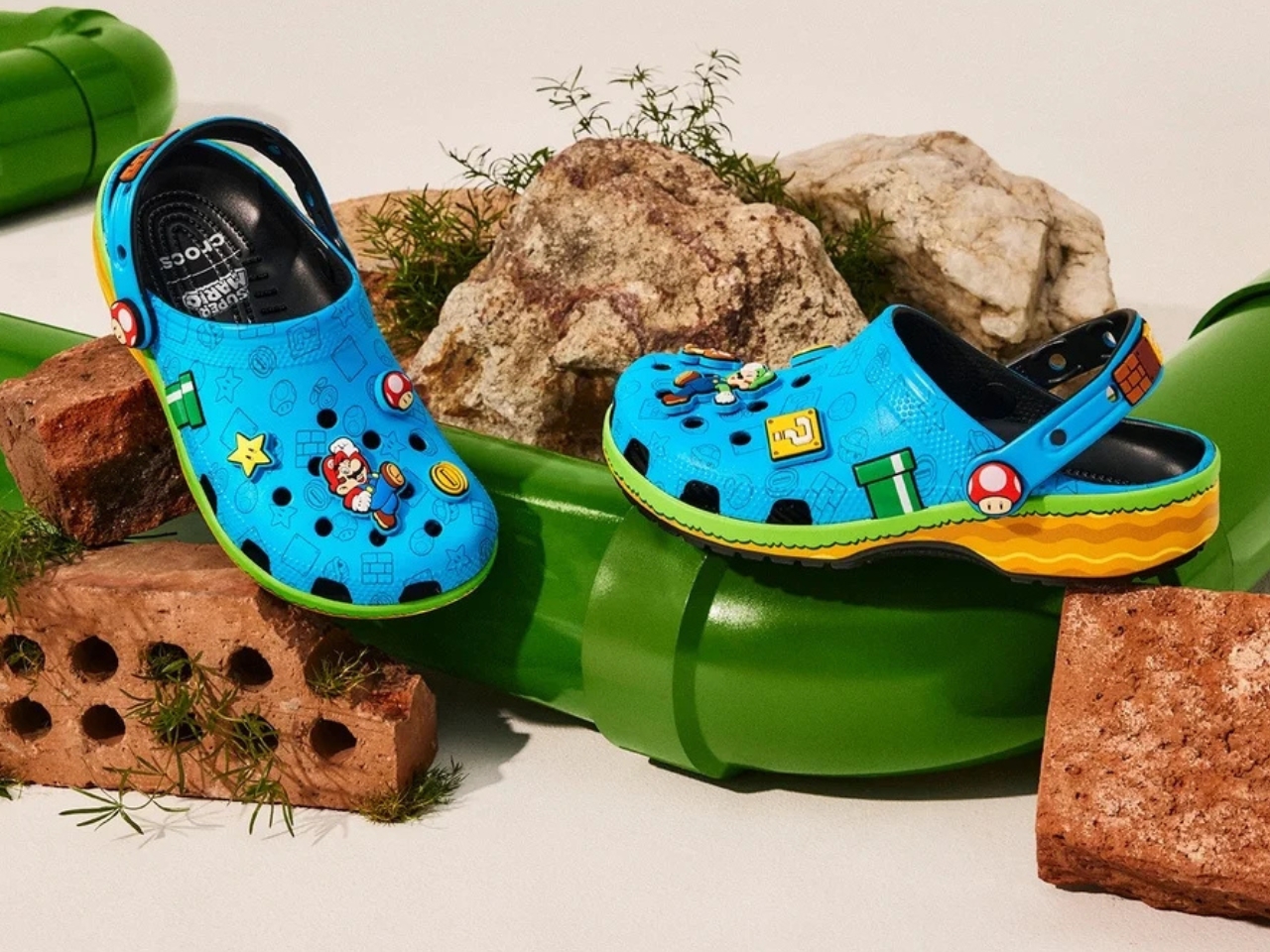

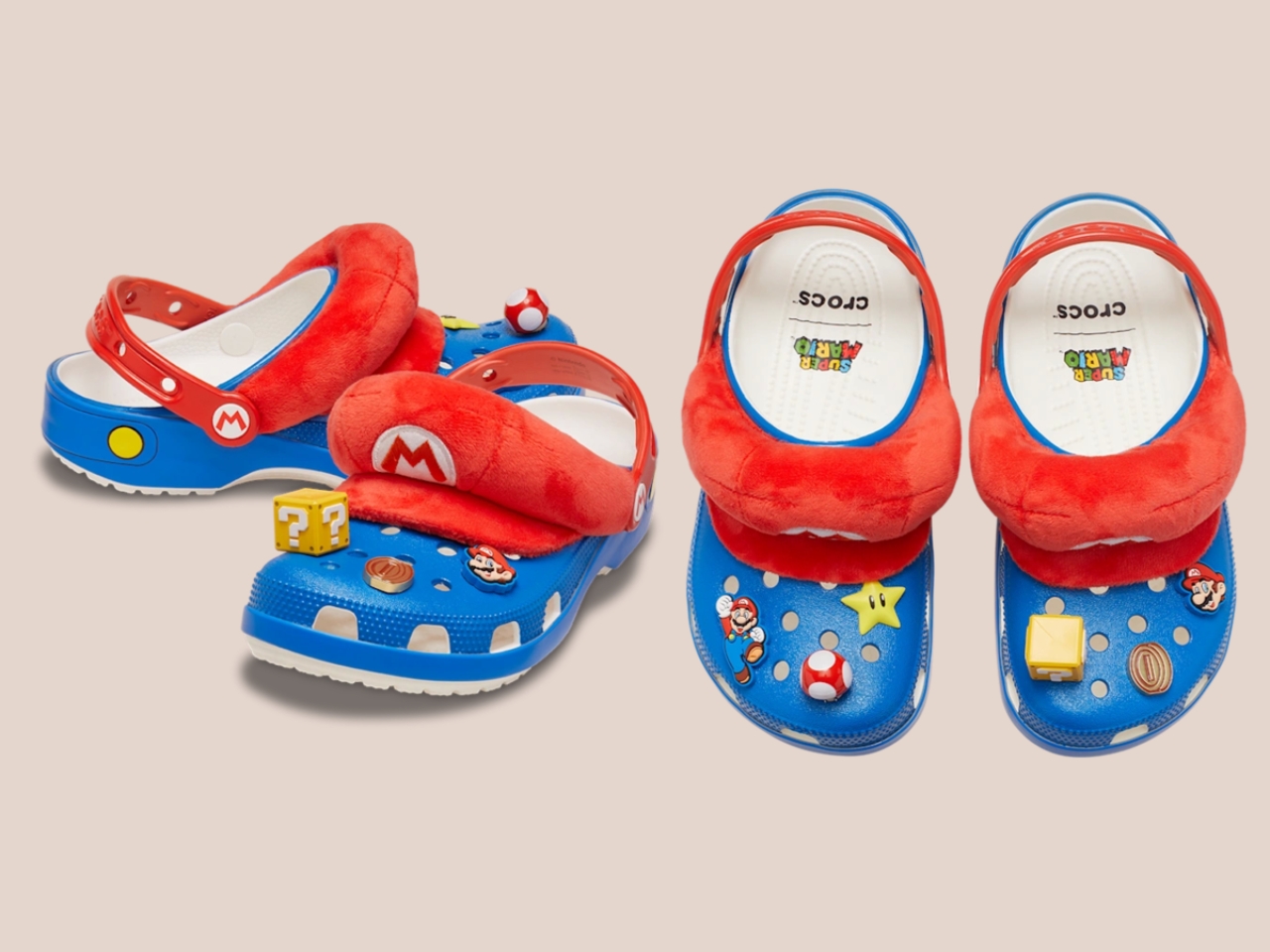
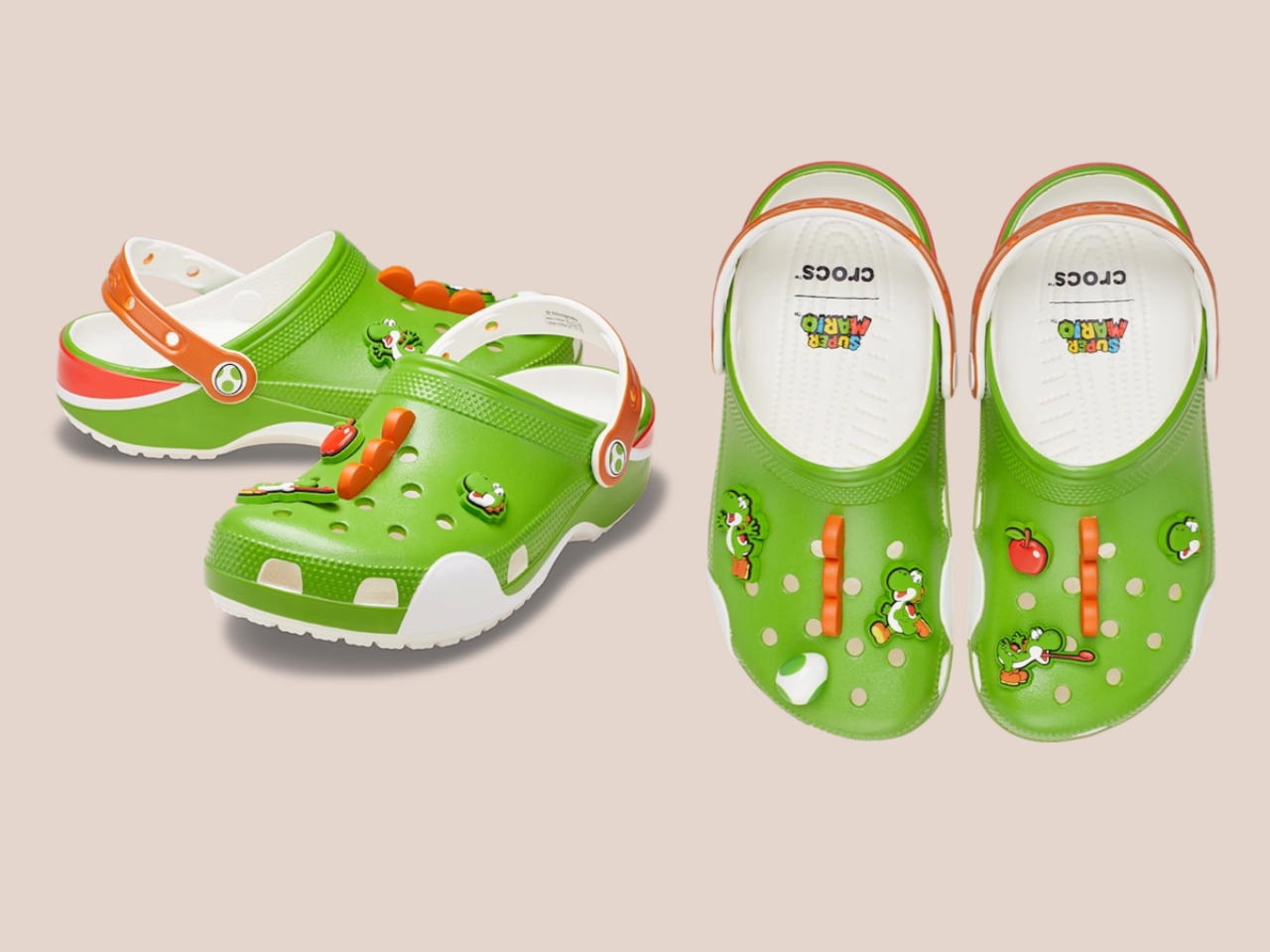
Five styles make up the collection, each built around a specific character, and the level of detail is more than you’d expect from a licensed shoe drop. The Mario clog sits on a blue foam base with a plush version of Mario’s signature red cap attached directly to the upper — playful without being costume-y. The Yoshi style goes full green with PVC fins running along the upper, which somehow manages to feel sculptural rather than silly. Then there’s the Bowser clog: dark green foam, seven PVC spikes on the upper, and five more replacing Jibbitz on the strap. Bowser would absolutely approve.
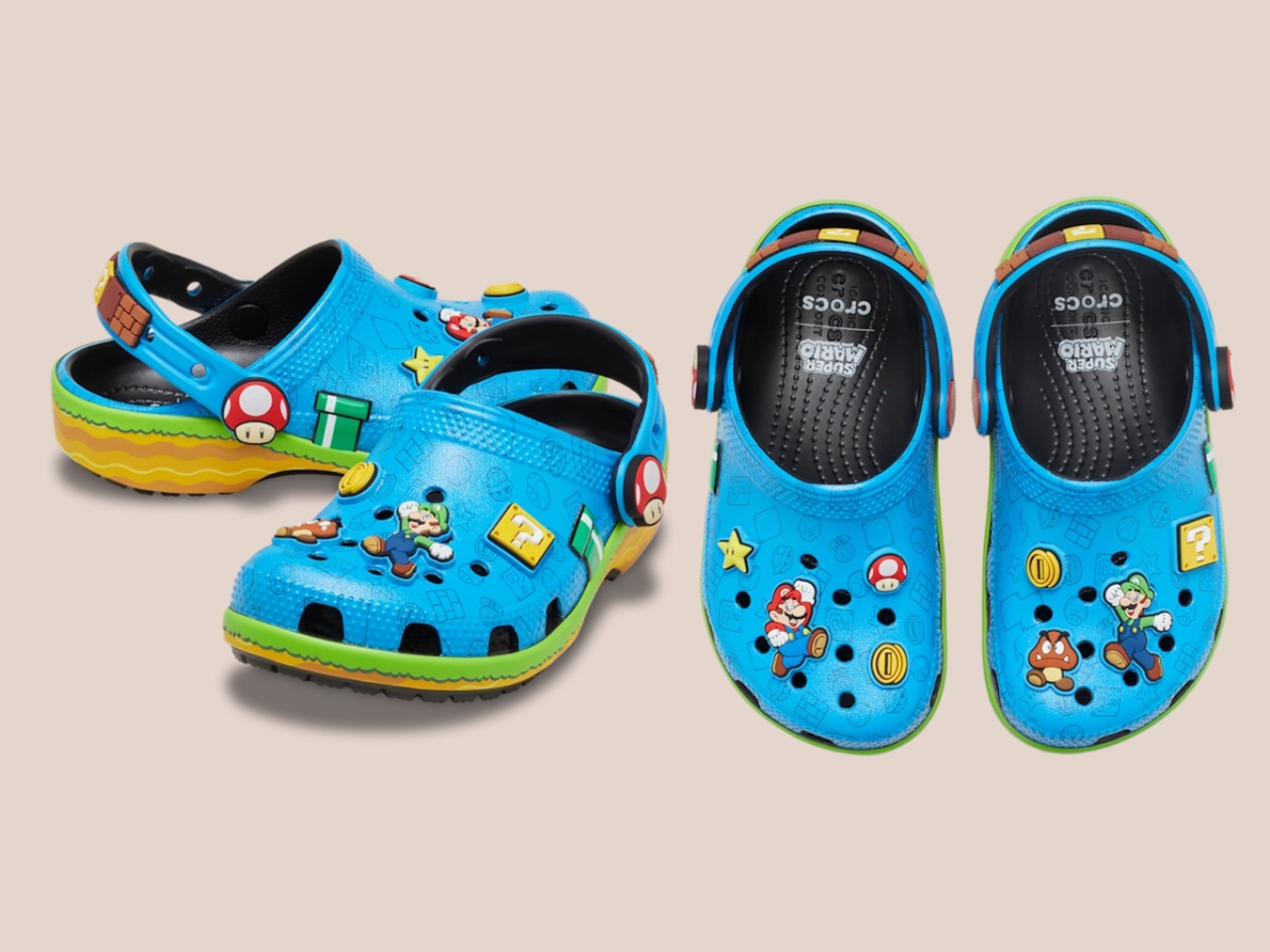



Princess Peach gets the most elevated treatment, with a platform sole finished in pink glitter and seven exclusive Jibbitz charms. It reads more fashion than novelty, which is a genuine design achievement for a shoe that has a question block on it. Rounding out the group is the Core Classic Clog, which pulls from the game’s broader iconography — coins, stars, turtle shells — and comes with eight Jibbitz charms and an option in kids’ sizing. The right call.
Pricing runs from $55 to $90, which lands comfortably in impulse-buy territory for any adult who will absolutely tell themselves they’re buying it for their kid.
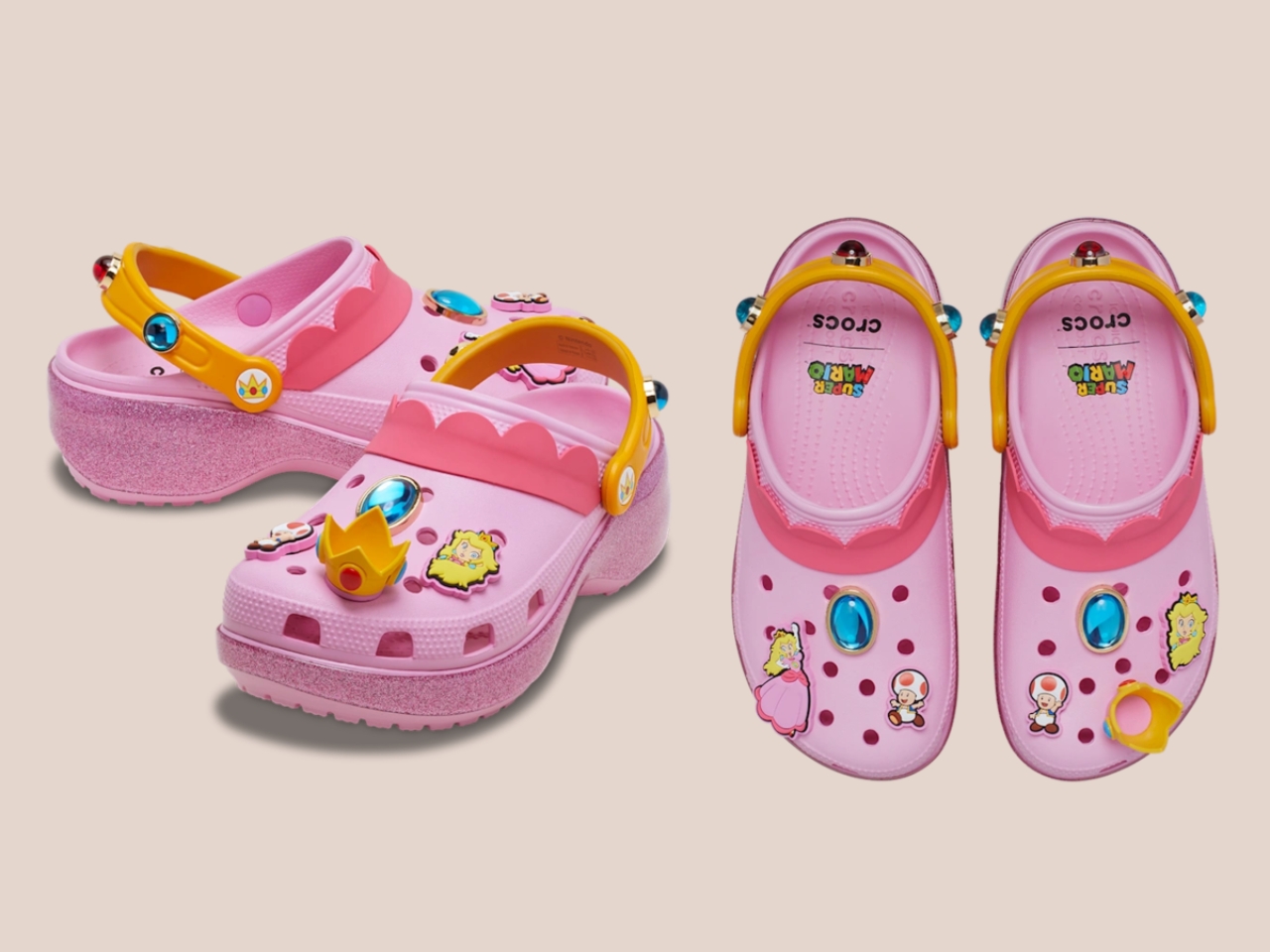
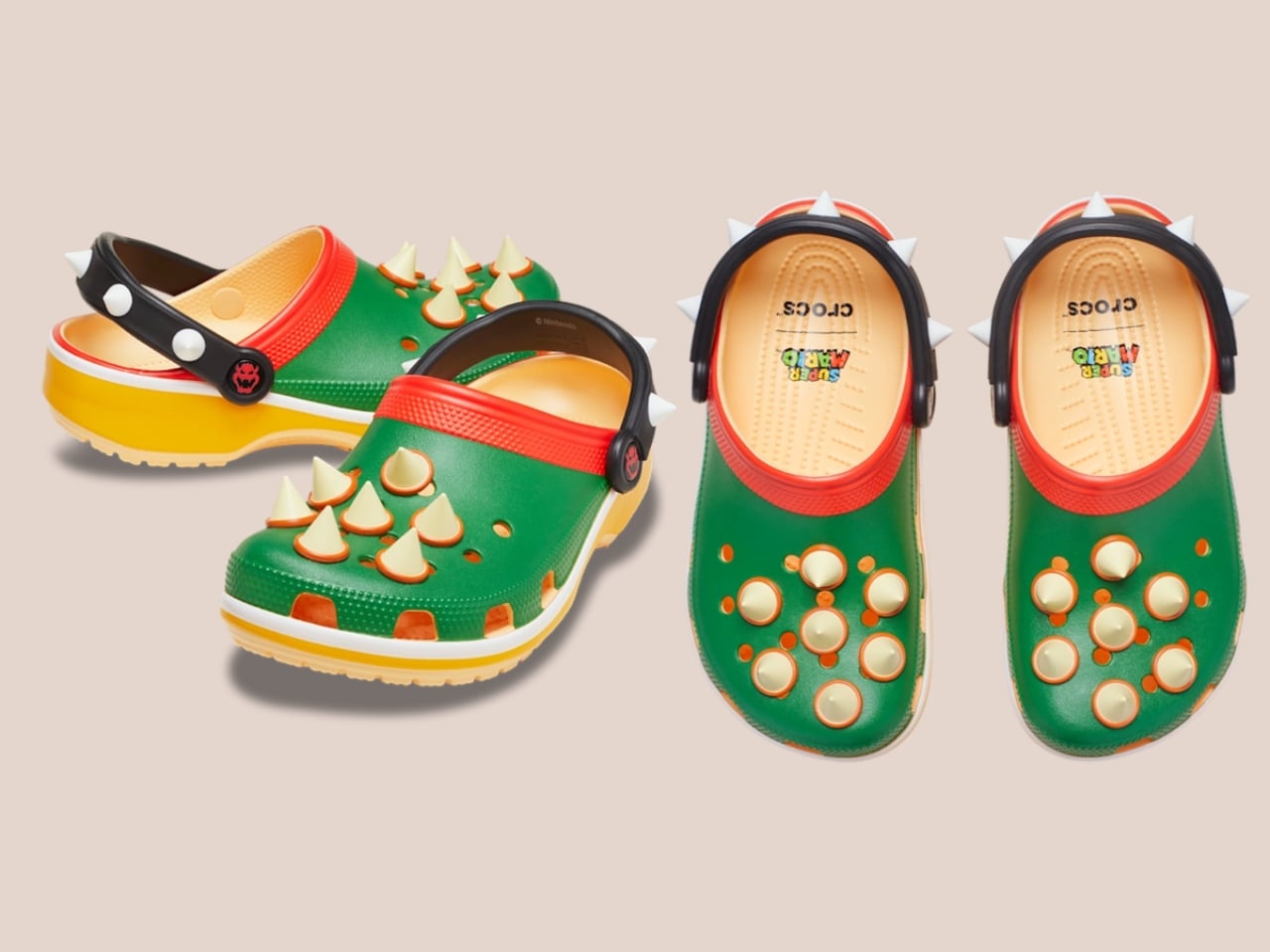


What makes this feel considered rather than purely commercial is how naturally the characters map onto what Crocs already does. Crocs has never been a brand that does subtle. It’s loud, comfortable, and entirely unapologetic about existing. Nintendo’s Super Mario roster operates the same way. These are characters drawn in primary colours with maximum personality and zero pretension. They don’t need your endorsement. So when Bowser turns up as a spike-covered clog, it doesn’t feel like a brand sticking a logo on a product. It feels like the character found the right medium.
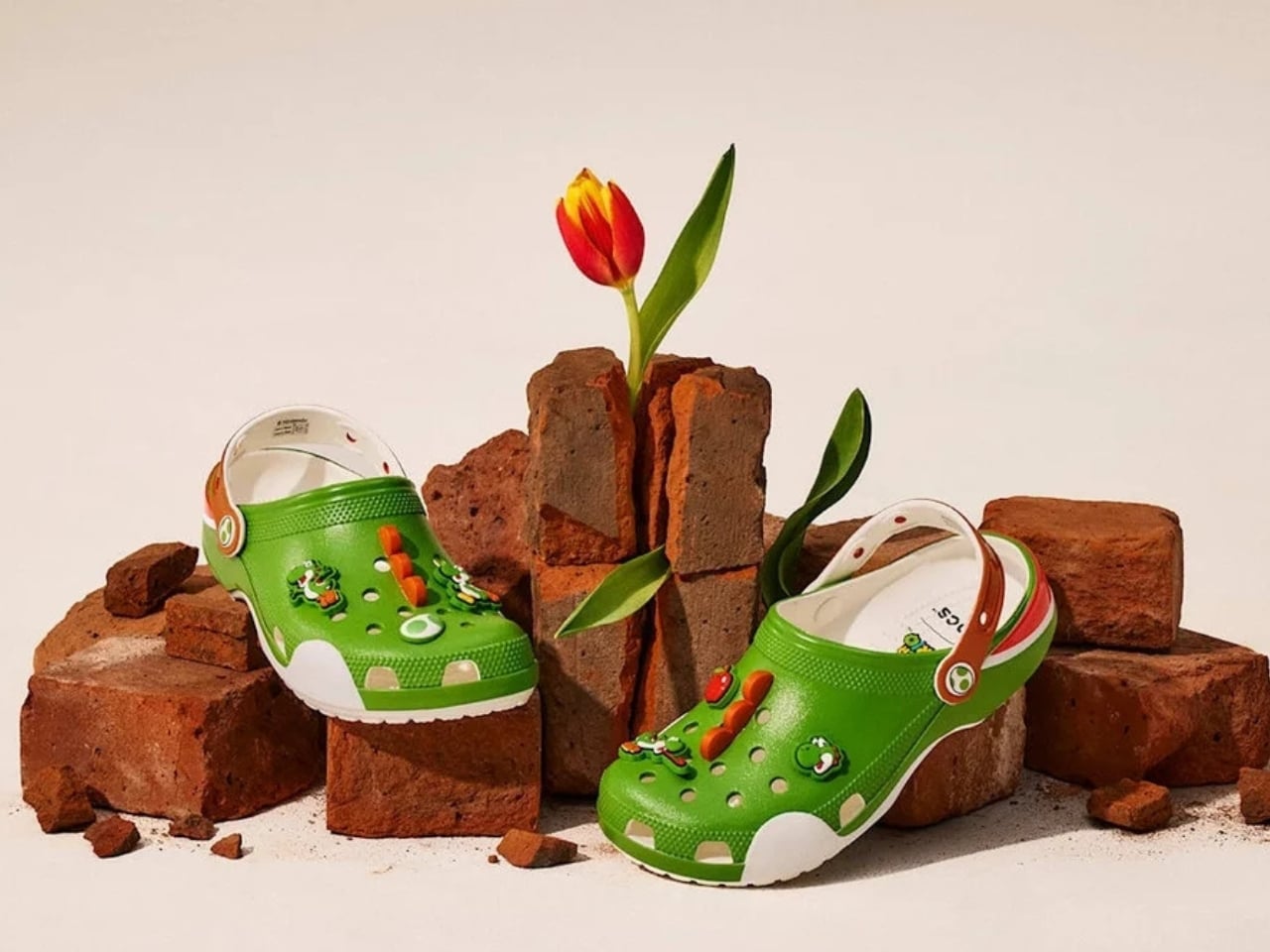

The Jibbitz element is doing quiet but important work across the collection. Each pair ships with six to eight exclusive co-branded charms, which means every clog doubles as an entry point into Crocs’ customisation ecosystem. Crocs has always understood that Jibbitz turn a shoe into something personal, closer to a wearable mood board than just footwear. Giving Super Mario fans a way into that system is smart, and it means the collection will live well past its launch date.
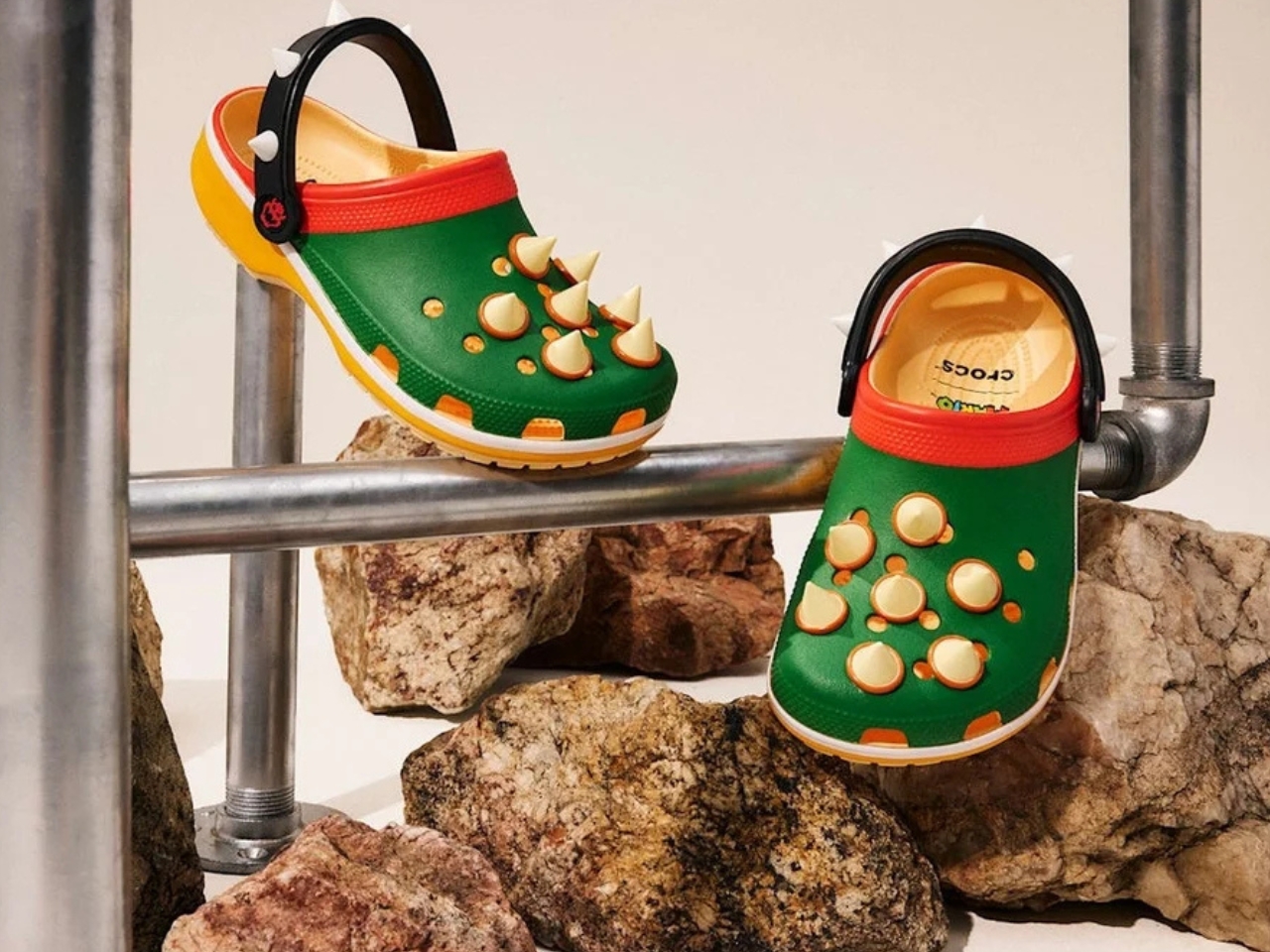

I’ll be straight: I did not expect to be genuinely interested in a pair of foam shoes shaped like Yoshi. I expected novelty for children and maybe a few nostalgic adults grabbing one on a whim. What the collection actually delivers is a thoughtful translation of game design into product design — character-specific textures, shapes, and details that suggest someone actually played the games and asked what it means for a shoe to feel like a character, not just look like one.
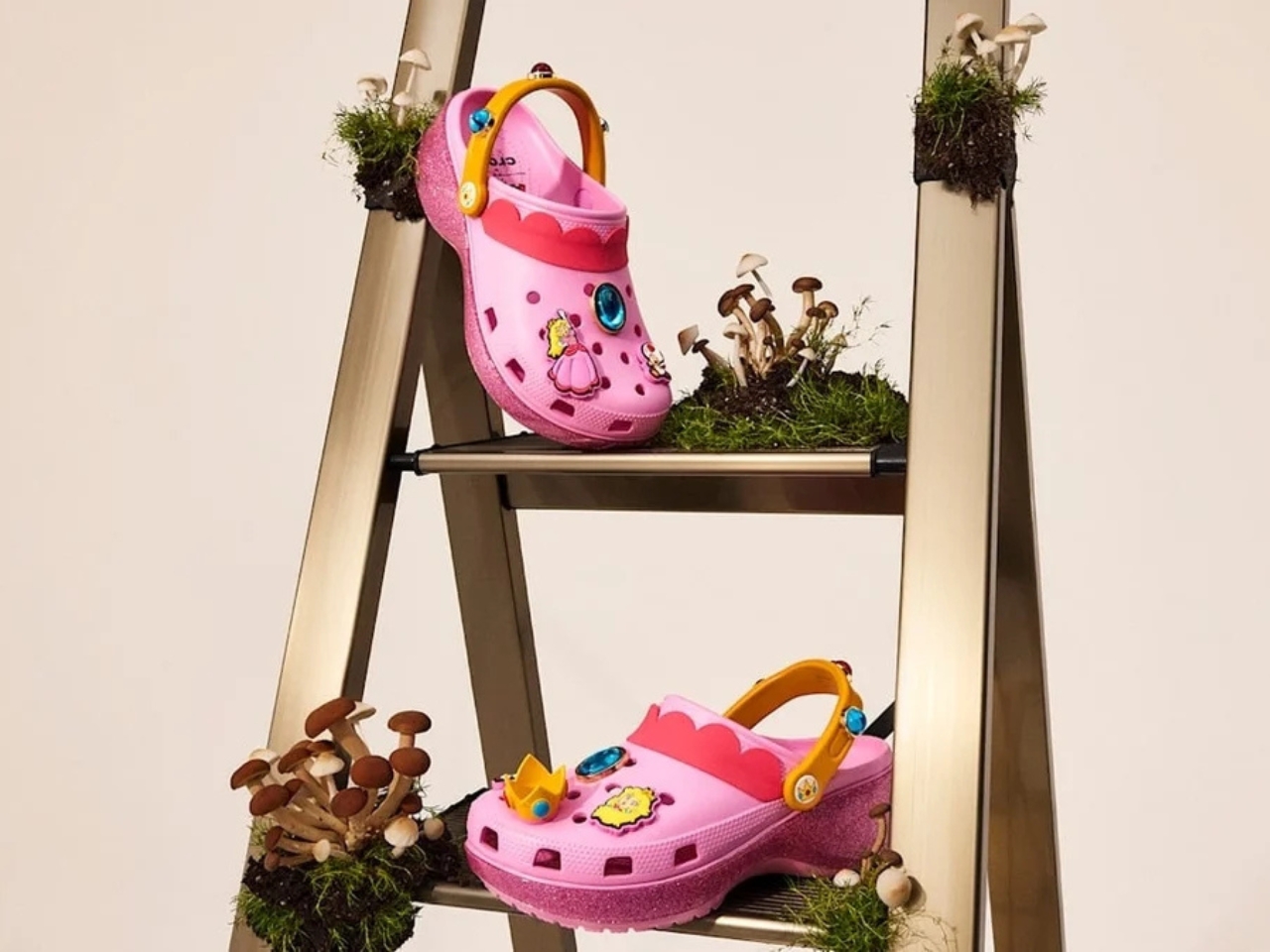

The Crocs x Super Mario collection is available from July 15 on crocs.com, the Nintendo Store, and select retailers globally. Your feet are going to the Mushroom Kingdom.

The post Crocs Just Entered the Mushroom Kingdom and We’re Into It first appeared on Yanko Design.
Apple said ‘AI’ exactly 28 times at WWDC 2026. Google mentioned it nearly 100 times at I/O.
Monday, June 8th, 2026 09:30 pmBy the end of this year’s big tech keynotes, one comparison stood out more than any product demo. Apple said “AI” 28 times at WWDC 2026. Google said it nearly 100 times at I/O 2026. Same industry, same race, same obsession, but two very different instincts about how to sell the next phase of computing.
Google’s keynote reflected the current rhythm of the AI industry, loud, relentless, and eager to stamp the term onto everything in sight. Apple’s presentation moved differently. It kept circling back to what people could actually do with the technology, how private it would be, and where it would fit into everyday routines. That softer framing may frustrate people who want Apple to move faster and compete harder. It may also be exactly why Apple’s pitch feels easier to absorb at a moment when audiences are already saturated with AI promises.

AI fatigue is real, and it has been building for a while. After years of keynotes, product launches, and press releases leading with the same two letters, the word has started to lose its grip on audiences. What once signaled breakthrough capability now signals marketing effort. When a company says “AI” 100 times in a single presentation, the listener stops hearing a technology and starts hearing a strategy. The signal becomes noise, and somewhere in that noise, the actual products get harder to see.
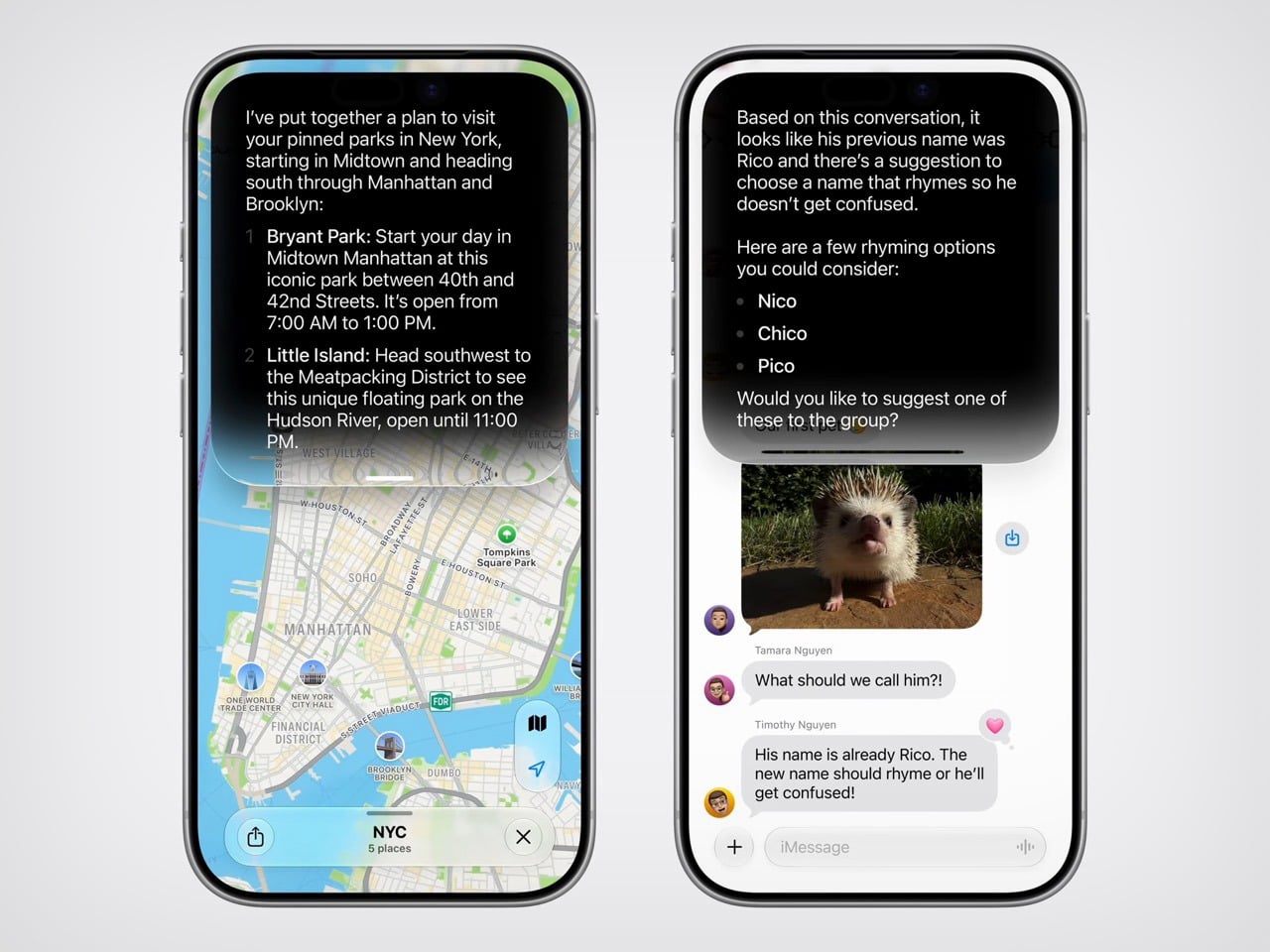
Apple’s approach at WWDC 2026 worked around that problem by reframing the conversation entirely. Instead of leading with technology, it led with moments. Siri finding a friend’s new address buried in a weeks-old message thread. A photo being reframed after the fact, as if you had stepped to the right before pressing the shutter. A restaurant bill split with Apple Cash by pointing a camera at it. These are small things, but they are the kind of small things that people actually think about during their day. Anchoring the keynote to those moments gave the technology a human scale that raw AI talk rarely achieves.
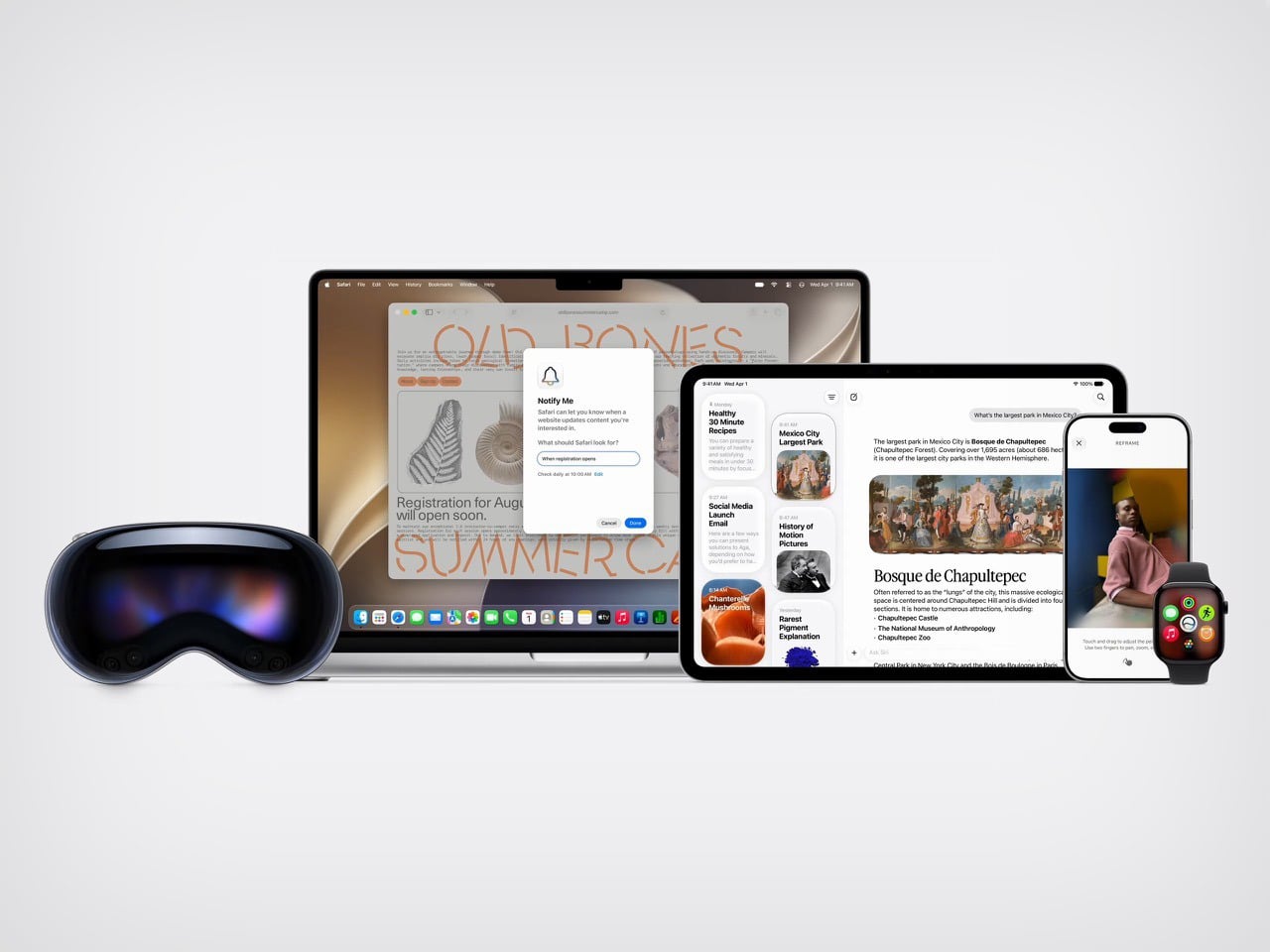
The branding reflects the same thinking. Apple calls it “Apple Intelligence,” a label that keeps the company name front and center while quietly sidestepping the overcrowded AI conversation. It is a deliberate choice, and it shows. Google’s keynote was structured around the technology itself, its power, its speed, its range. Apple’s keynote was structured around the people using it. That difference in framing shapes how audiences receive the same underlying capability, and Apple’s version is considerably easier to trust.
Privacy played a central role in building that trust. Apple returned to on-device processing and Private Cloud Compute repeatedly throughout WWDC, not as a footnote but as a feature. At a time when public concern about how AI companies handle personal data is growing steadily, that emphasis lands differently than it might have a few years ago. Google builds powerful models and serves them at enormous scale. Apple builds careful models and makes a point of telling you where your data goes and where it stays. For a meaningful portion of consumers, that distinction matters more than benchmark scores.

None of this means Apple is winning the AI race on capability. Google’s models are more powerful, more publicly accessible, and more deeply woven into the daily workflows of people around the world. Gemini’s reach across Search, Gmail, YouTube, and Android gives Google a distribution advantage that Apple’s ecosystem, for all its loyalty, cannot easily match. If the competition were judged purely on technical ambition and model performance, Google’s 100 mentions would feel earned.
But technology keynotes are not judged purely on technical ambition. They are judged on how they make audiences feel, what they make people want, and whether they leave the room energised or overwhelmed. On those terms, Apple’s 28 mentions of “AI” accomplished something that Google’s near-100 did not. They kept the word rare enough to mean something. Every time Apple said it, there was a feature attached, a privacy assurance nearby, and a use case grounded in daily life. The word carried weight because it was not being used to fill space.


The larger irony is that Apple may be the company best positioned to benefit from a backlash it did not entirely create. Google, Microsoft, Meta, and others have spent years flooding the conversation with AI language, and the fatigue that has followed is a byproduct of their own enthusiasm. Apple watched, built quietly, and showed up at WWDC 2026 with a keynote that treated restraint as a product decision. Whether that restraint reflects genuine strategic confidence or simply a capability gap dressed up in good marketing is the question the next few years will answer. For now, 28 versus 100 tells a story that Apple’s communications team could not have scripted better.

The post Apple said ‘AI’ exactly 28 times at WWDC 2026. Google mentioned it nearly 100 times at I/O. first appeared on Yanko Design.